Reading the Combinatorial Histone Language
- PMID: 26675328
- PMCID: PMC10127456
- DOI: 10.1021/acschembio.5b00864
Reading the Combinatorial Histone Language
Abstract
Histones are subject to frequent combinatorial post-translational modifications (PTMs), forming a complex chemical "language" that is interpreted by PTM-specific histone-interacting protein modules (reader domains). These specific interactions are thought to instruct gene expression and downstream biological functions. While the majority of studies have focused on individual modifications, our current understanding of the combinatorial PTM patterns on histones is starting to emerge, benefiting from the convergence of multiple technologies. Here, we review the key technical advances and progress on discovery and characterization of combinatorial histone PTM patterns. We focus on the interactions between reader domains and combinatorial PTMs, which is essential for understanding the mechanism and biological meaning of establishing and interpreting information embedded in histone PTM patterns.
Figures
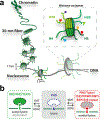



References
-
- Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA, Science 184, 868–871. - PubMed
-
- Kornberg RD, and Klug A. (1981) The nucleosome, Scientific American 244, 52–64. - PubMed
-
- Kornberg RD, and Lorch Y. (1999) Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome, Cell 98, 285–294. - PubMed
-
- Li G, and Zhu P. (2015) Structure and organization of chromatin fiber in the nucleus, FEBS letters. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

