Identification of potential drug targets by subtractive genome analysis of Escherichia coli O157:H7: an in silico approach
- PMID: 26677339
- PMCID: PMC4677596
- DOI: 10.2147/AABC.S88522
Identification of potential drug targets by subtractive genome analysis of Escherichia coli O157:H7: an in silico approach
Abstract
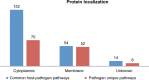
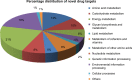
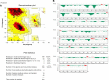
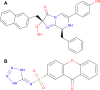
Bacterial enteric infections resulting in diarrhea, dysentery, or enteric fever constitute a huge public health problem, with more than a billion episodes of disease annually in developing and developed countries. In this study, the deadly agent of hemorrhagic diarrhea and hemolytic uremic syndrome, Escherichia coli O157:H7 was investigated with extensive computational approaches aimed at identifying novel and broad-spectrum antibiotic targets. A systematic in silico workflow consisting of comparative genomics, metabolic pathways analysis, and additional drug prioritizing parameters was used to identify novel drug targets that were essential for the pathogen's survival but absent in its human host. Comparative genomic analysis of Kyoto Encyclopedia of Genes and Genomes annotated metabolic pathways identified 350 putative target proteins in E. coli O157:H7 which showed no similarity to human proteins. Further bio-informatic approaches including prediction of subcellular localization, calculation of molecular weight, and web-based investigation of 3D structural characteristics greatly aided in filtering the potential drug targets from 350 to 120. Ultimately, 44 non-homologous essential proteins of E. coli O157:H7 were prioritized and proved to have the eligibility to become novel broad-spectrum antibiotic targets and DNA polymerase III alpha (dnaE) was the top-ranked among these targets. Moreover, druggability of each of the identified drug targets was evaluated by the DrugBank database. In addition, 3D structure of the dnaE was modeled and explored further for in silico docking with ligands having potential druggability. Finally, we confirmed that the compounds N-coeleneterazine and N-(1,4-dihydro-5H-tetrazol-5-ylidene)-9-oxo-9H-xanthene-2-sulfon-amide were the most suitable ligands of dnaE and hence proposed as the potential inhibitors of this target protein. The results of this study could facilitate the discovery and release of new and effective drugs against E. coli O157:H7 and other deadly human bacterial pathogens.
Keywords: DNA polymerase III alpha; E. coli O157:H7; KEGG metabolic pathways; homology modeling; novel and broad-spectrum antibiotic targets.
Figures




Similar articles
-
Relative nephroprotection during Escherichia coli O157:H7 infections: association with intravenous volume expansion.Pediatrics. 2005 Jun;115(6):e673-80. doi: 10.1542/peds.2004-2236. Pediatrics. 2005. PMID: 15930195
-
Rapid identification of protein biomarkers of Escherichia coli O157:H7 by matrix-assisted laser desorption ionization-time-of-flight-time-of-flight mass spectrometry and top-down proteomics.Anal Chem. 2010 Apr 1;82(7):2717-25. doi: 10.1021/ac902455d. Anal Chem. 2010. PMID: 20232878
-
Illnesses associated with Escherichia coli O157:H7 infections. A broad clinical spectrum.Ann Intern Med. 1988 Nov 1;109(9):705-12. doi: 10.7326/0003-4819-109-9-705. Ann Intern Med. 1988. PMID: 3056169 Review.
-
From In silico Protein Epitope Density Prediction to Testing Escherichia coli O157:H7 Vaccine Candidates in a Murine Model of Colonization.Front Cell Infect Microbiol. 2016 Aug 30;6:94. doi: 10.3389/fcimb.2016.00094. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27625996 Free PMC article.
-
Escherichia coli O157:H7 infection in humans.Ann Intern Med. 1995 Nov 1;123(9):698-714. doi: 10.7326/0003-4819-123-9-199511010-00009. Ann Intern Med. 1995. PMID: 7574226 Review.
Cited by
-
An integrative, multi-omics approach towards the prioritization of Klebsiella pneumoniae drug targets.Sci Rep. 2018 Jul 17;8(1):10755. doi: 10.1038/s41598-018-28916-7. Sci Rep. 2018. PMID: 30018343 Free PMC article.
-
In Silico Screening and Analysis of Broad-Spectrum Molecular Targets and Lead Compounds for Diarrhea Therapy.Bioinform Biol Insights. 2019 Oct 24;13:1177932219884297. doi: 10.1177/1177932219884297. eCollection 2019. Bioinform Biol Insights. 2019. PMID: 31695343 Free PMC article.
-
Study of intra-inter species protein-protein interactions for potential drug targets identification and subsequent drug design for Escherichia coli O104:H4 C277-11.In Silico Pharmacol. 2016 Dec;5(1):1. doi: 10.1007/s40203-017-0021-5. Epub 2017 Apr 11. In Silico Pharmacol. 2016. PMID: 28401513 Free PMC article.
-
Finding Potential Therapeutic Targets against Shigella flexneri through Proteome Exploration.Front Microbiol. 2016 Nov 22;7:1817. doi: 10.3389/fmicb.2016.01817. eCollection 2016. Front Microbiol. 2016. PMID: 27920755 Free PMC article.
-
Genome-Based Drug Target Identification in Human Pathogen Streptococcus gallolyticus.Front Genet. 2021 Mar 25;12:564056. doi: 10.3389/fgene.2021.564056. eCollection 2021. Front Genet. 2021. PMID: 33841489 Free PMC article.
References
-
- Kaper JB, Nataro JP, Mobley HL. Pathogenic Escherichia coli. Nat Rev Microbiol. 2004;2(2):123–140. - PubMed
-
- Law D. Virulence factors of Escherichia coli O157 and other Shiga toxin-producing E. coli. J Appl Microbiol. 2000;88(5):729–745. - PubMed
-
- Ostroff SM, Griffin PM, Tauxe RV, et al. A statewide outbreak of Escherichia coli O157:H7 infections in Washington State. Am J Epidemiol. 1990;132(2):239–247. - PubMed
-
- Griffin PM, Tauxe RV. The epidemiology of infections caused by Escherichia coli O157:H7, other enterohemorrhagic E. coli, and the associated hemolytic uremic syndrome. Epidemiol Rev. 1991;13:60–98. - PubMed
-
- Besser RE, Lett SM, Weber JT, et al. An outbreak of diarrhea and hemolytic uremic syndrome from Escherichia coli O157:H7 in fresh-pressed apple cider. JAMA. 1993;269(17):2217–2220. - PubMed
LinkOut - more resources
Full Text Sources

