Evidence of widespread natural recombination among field isolates of equine herpesvirus 4 but not among field isolates of equine herpesvirus 1
- PMID: 26691326
- PMCID: PMC5381393
- DOI: 10.1099/jgv.0.000378
Evidence of widespread natural recombination among field isolates of equine herpesvirus 4 but not among field isolates of equine herpesvirus 1
Abstract
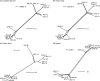
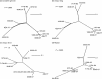
Recombination in alphaherpesviruses allows evolution to occur in viruses that have an otherwise stable DNA genome with a low rate of nucleotide substitution. High-throughput sequencing of complete viral genomes has recently allowed natural (field) recombination to be studied in a number of different alphaherpesviruses, however, such studies have not been applied to equine herpesvirus 1 (EHV-1) or equine herpesvirus 4 (EHV-4). These two equine alphaherpesviruses are genetically similar, but differ in their pathogenesis and epidemiology. Both cause economically significant disease in horse populations worldwide. This study used high-throughput sequencing to determine the full genome sequences of EHV-1 and EHV-4 isolates (11 and 14 isolates, respectively) from Australian or New Zealand horses. These sequences were then analysed and examined for evidence of recombination. Evidence of widespread recombination was detected in the genomes of the EHV-4 isolates. Only one potential recombination event was detected in the genomes of the EHV-1 isolates, even when the genomes from an additional 11 international EHV-1 isolates were analysed. The results from this study reveal another fundamental difference between the biology of EHV-1 and EHV-4. The results may also be used to help inform the future safe use of attenuated equine herpesvirus vaccines.
Figures


Similar articles
-
The full genome sequences of 8 equine herpesvirus type 4 isolates from horses in Japan.J Vet Med Sci. 2017 Jan 24;79(1):206-212. doi: 10.1292/jvms.16-0506. Epub 2016 Nov 14. J Vet Med Sci. 2017. PMID: 27840393 Free PMC article.
-
Comparing the genetic diversity of ORF30 of Australian isolates of 3 equid alphaherpesviruses.Vet Microbiol. 2014 Feb 21;169(1-2):50-7. doi: 10.1016/j.vetmic.2013.12.007. Epub 2013 Dec 25. Vet Microbiol. 2014. PMID: 24418044
-
Temporal detection of equine herpesvirus infections of a cohort of mares and their foals.Vet Microbiol. 2006 Sep 10;116(4):249-57. doi: 10.1016/j.vetmic.2006.05.002. Epub 2006 Jun 13. Vet Microbiol. 2006. PMID: 16774810
-
Equine herpesviruses type 1 (EHV-1) and 4 (EHV-4)--masters of co-evolution and a constant threat to equids and beyond.Vet Microbiol. 2013 Nov 29;167(1-2):123-34. doi: 10.1016/j.vetmic.2013.06.018. Epub 2013 Jul 6. Vet Microbiol. 2013. PMID: 23890672 Review.
-
A review of equid herpesvirus 1 for the veterinary practitioner. Part A: clinical presentation, diagnosis and treatment.N Z Vet J. 2014 Jul;62(4):171-8. doi: 10.1080/00480169.2014.899945. N Z Vet J. 2014. PMID: 24597778 Review.
Cited by
-
Equid alphaherpesvirus 1 from Italian Horses: Evaluation of the Variability of the ORF30, ORF33, ORF34 and ORF68 Genes.Viruses. 2019 Sep 13;11(9):851. doi: 10.3390/v11090851. Viruses. 2019. PMID: 31540321 Free PMC article.
-
Outbreak of equine herpesvirus 4 (EHV-4) in Denmark: tracing patient zero and viral characterization.BMC Vet Res. 2024 Jul 3;20(1):287. doi: 10.1186/s12917-024-04149-x. BMC Vet Res. 2024. PMID: 38961400 Free PMC article.
-
Genomic, Recombinational and Phylogenetic Characterization of Global Feline Herpesvirus 1 Isolates.Virology. 2018 May;518:385-397. doi: 10.1016/j.virol.2018.03.018. Epub 2018 Mar 30. Virology. 2018. PMID: 29605685 Free PMC article.
-
The full genome sequences of 8 equine herpesvirus type 4 isolates from horses in Japan.J Vet Med Sci. 2017 Jan 24;79(1):206-212. doi: 10.1292/jvms.16-0506. Epub 2016 Nov 14. J Vet Med Sci. 2017. PMID: 27840393 Free PMC article.
-
Molecular Characterisation of Equine Herpesvirus 1 Isolates from Cases of Abortion, Respiratory and Neurological Disease in Ireland between 1990 and 2017.Pathogens. 2019 Jan 15;8(1):7. doi: 10.3390/pathogens8010007. Pathogens. 2019. PMID: 30650561 Free PMC article.
References
-
- Allen G., Kydd J., Slater J. (2004). Equid herpesvirus 1 and equid herpesvirus 4 infections. In Infectious Diseases of Livestock, pp. 829–859. Edited by Coetzer J., Tustin R.Newmarket: Oxford University Press.
-
- Bagust T. J., Pascoe R. R. (1968). Isolation of equine rhinopnemonitis virus from acute respiratory disease in a horse in queensland Vet J 44296.
-
- Bagust T. J., Pascoe R. R. (1970). Studies on equine herpesviruses. 1. Characterisation of a strain of equine rhinopneumonitis virus isolated in Queensland Aust Vet J 46421–427. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

