Functional Divergence of APETALA1 and FRUITFULL is due to Changes in both Regulation and Coding Sequence
- PMID: 26697035
- PMCID: PMC4667048
- DOI: 10.3389/fpls.2015.01076
Functional Divergence of APETALA1 and FRUITFULL is due to Changes in both Regulation and Coding Sequence
Abstract
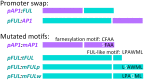
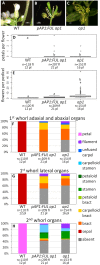
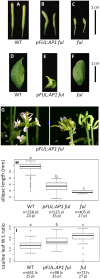
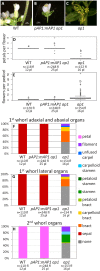
Gene duplications are prevalent in plants, and functional divergence subsequent to duplication may be linked with the occurrence of novel phenotypes in plant evolution. Here, we examine the functional divergence of Arabidopsis thaliana APETALA1 (AP1) and FRUITFULL (FUL), which arose via a duplication correlated with the origin of the core eudicots. Both AP1 and FUL play a role in floral meristem identity, but AP1 is required for the formation of sepals and petals whereas FUL is involved in cauline leaf and fruit development. AP1 and FUL are expressed in mutually exclusive domains but also differ in sequence, with unique conserved motifs in the C-terminal domains of the proteins that suggest functional differentiation. To determine whether the functional divergence of AP1 and FUL is due to changes in regulation or changes in coding sequence, we performed promoter swap experiments, in which FUL was expressed in the AP1 domain in the ap1 mutant and vice versa. Our results show that FUL can partially substitute for AP1, and AP1 can partially substitute for FUL; thus, the functional divergence between AP1 and FUL is due to changes in both regulation and coding sequence. We also mutated AP1 and FUL conserved motifs to determine if they are required for protein function and tested the ability of these mutated proteins to interact in yeast with known partners. We found that these motifs appear to play at best a minor role in protein function and dimerization capability, despite being strongly conserved. Our results suggest that the functional differentiation of these two paralogous key transcriptional regulators involves both differences in regulation and in sequence; however, sequence changes in the form of unique conserved motifs do not explain the differences observed.
Keywords: APETALA1; FRUITFULL; MADS box genes; conserved protein motifs; functional divergence; gene duplication.
Figures

Similar articles
-
Poppy APETALA1/FRUITFULL orthologs control flowering time, branching, perianth identity, and fruit development.Plant Physiol. 2012 Apr;158(4):1685-704. doi: 10.1104/pp.111.192104. Epub 2012 Jan 27. Plant Physiol. 2012. PMID: 22286183 Free PMC article.
-
Assessing duplication and loss of APETALA1/FRUITFULL homologs in Ranunculales.Front Plant Sci. 2013 Sep 17;4:358. doi: 10.3389/fpls.2013.00358. eCollection 2013. Front Plant Sci. 2013. PMID: 24062757 Free PMC article.
-
Duplication of AP1 within the Spinacia oleracea L. AP1/FUL clade is followed by rapid amino acid and regulatory evolution.Planta. 2009 Feb;229(3):507-21. doi: 10.1007/s00425-008-0851-9. Epub 2008 Nov 13. Planta. 2009. PMID: 19005675
-
Functional Conservation and Divergence of Five AP1/FUL-like Genes in Marigold (Tagetes erecta L.).Genes (Basel). 2021 Dec 17;12(12):2011. doi: 10.3390/genes12122011. Genes (Basel). 2021. PMID: 34946960 Free PMC article.
-
The Aquilegia FRUITFULL-like genes play key roles in leaf morphogenesis and inflorescence development.Plant J. 2013 Apr;74(2):197-212. doi: 10.1111/tpj.12113. Epub 2013 Mar 7. Plant J. 2013. PMID: 23294330
Cited by
-
Utilizing MIKC-type MADS-box protein SOC1 for yield potential enhancement in maize.Plant Cell Rep. 2021 Sep;40(9):1679-1693. doi: 10.1007/s00299-021-02722-4. Epub 2021 Jun 6. Plant Cell Rep. 2021. PMID: 34091722 Free PMC article.
-
RcAP1, a Homolog of APETALA1, is Associated with Flower Bud Differentiation and Floral Organ Morphogenesis in Rosa chinensis.Int J Mol Sci. 2019 Jul 20;20(14):3557. doi: 10.3390/ijms20143557. Int J Mol Sci. 2019. PMID: 31330828 Free PMC article.
-
Genome-wide identification, interaction of the MADS-box proteins in Zanthoxylum armatum and functional characterization of ZaMADS80 in floral development.Front Plant Sci. 2022 Nov 25;13:1038828. doi: 10.3389/fpls.2022.1038828. eCollection 2022. Front Plant Sci. 2022. PMID: 36507394 Free PMC article.
-
Divergent Functional Diversification Patterns in the SEP/AGL6/AP1 MADS-Box Transcription Factor Superclade.Plant Cell. 2019 Dec;31(12):3033-3056. doi: 10.1105/tpc.19.00162. Epub 2019 Oct 7. Plant Cell. 2019. PMID: 31591161 Free PMC article.
-
Expression patterns of Passiflora edulis APETALA1/FRUITFULL homologues shed light onto tendril and corona identities.Evodevo. 2017 Feb 2;8:3. doi: 10.1186/s13227-017-0066-x. eCollection 2017. Evodevo. 2017. PMID: 28174623 Free PMC article.
References
-
- Berbel A., Navarro C., Ferrándiz C., Cañas L. A., Madueño F., Beltrán J.-P. (2001). Analysis of PEAM4, the pea AP1 functional homologue, supports a model for AP1-like genes controlling both floral meristem and floral organ identity in different plant species. Plant J. 25 441–451. 10.1046/j.1365-313x.2001.00974.x - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

