Three-Dimensional Gastrointestinal Organoid Culture in Combination with Nerves or Fibroblasts: A Method to Characterize the Gastrointestinal Stem Cell Niche
- PMID: 26697073
- PMCID: PMC4677245
- DOI: 10.1155/2016/3710836
Three-Dimensional Gastrointestinal Organoid Culture in Combination with Nerves or Fibroblasts: A Method to Characterize the Gastrointestinal Stem Cell Niche
Abstract
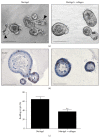
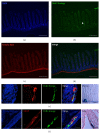
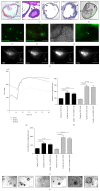
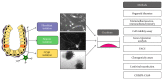
The gastrointestinal epithelium is characterized by a high turnover of cells and intestinal stem cells predominantly reside at the bottom of crypts and their progeny serve to maintain normal intestinal homeostasis. Accumulating evidence demonstrates the pivotal role of a niche surrounding intestinal stem cells in crypts, which consists of cellular and soluble components and creates an environment constantly influencing the fate of stem cells. Here we describe different 3D culture systems to culture gastrointestinal epithelium that should enable us to study the stem cell niche in vitro in the future: organoid culture and multilayered systems such as organotypic cell culture and culture of intestinal tissue fragments ex vivo. These methods mimic the in vivo situation in vitro by creating 3D culture conditions that reflect the physiological situation of intestinal crypts. Modifications of the composition of the culture media as well as coculturing epithelial organoids with previously described cellular components such as myofibroblasts, collagen, and neurons show the impact of the methods applied to investigate niche interactions in vitro. We further present a novel method to isolate labeled nerves from the enteric nervous system using Dclk1-CreGFP mice.
Figures



Similar articles
-
The Organoid Reconstitution Assay (ORA) for the Functional Analysis of Intestinal Stem and Niche Cells.J Vis Exp. 2017 Nov 20;(129):56329. doi: 10.3791/56329. J Vis Exp. 2017. PMID: 29286377 Free PMC article.
-
Isolation and In Vitro Culture of Human Gut Progenitor Cells.Methods Mol Biol. 2019;2029:49-62. doi: 10.1007/978-1-4939-9631-5_5. Methods Mol Biol. 2019. PMID: 31273733
-
Ex vivo culture of the intestinal epithelium: strategies and applications.Gut. 2014 Aug;63(8):1345-54. doi: 10.1136/gutjnl-2014-307204. Epub 2014 May 19. Gut. 2014. PMID: 24841573 Review.
-
Advanced three-dimensional culture of equine intestinal epithelial stem cells.Equine Vet J. 2018 Mar;50(2):241-248. doi: 10.1111/evj.12734. Epub 2017 Sep 6. Equine Vet J. 2018. PMID: 28792626 Free PMC article.
-
Mini-gut organoids: reconstitution of the stem cell niche.Annu Rev Cell Dev Biol. 2015;31:269-89. doi: 10.1146/annurev-cellbio-100814-125218. Epub 2015 Oct 2. Annu Rev Cell Dev Biol. 2015. PMID: 26436704 Review.
Cited by
-
Murine colon organoids as a novel model to study Trypanosoma cruzi infection and interactions with the intestinal epithelium.Front Cell Infect Microbiol. 2023 Mar 9;13:1082524. doi: 10.3389/fcimb.2023.1082524. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 36968103 Free PMC article.
-
Characterizing caspase-1 involvement during esophageal disease progression.Cancer Immunol Immunother. 2020 Dec;69(12):2635-2649. doi: 10.1007/s00262-020-02650-4. Epub 2020 Jul 1. Cancer Immunol Immunother. 2020. PMID: 32613271 Free PMC article.
-
Cholinergic Activation of Primary Human Derived Intestinal Epithelium Does Not Ameliorate TNF-α Induced Injury.Cell Mol Bioeng. 2020 Jun 24;13(5):487-505. doi: 10.1007/s12195-020-00633-0. eCollection 2020 Oct. Cell Mol Bioeng. 2020. PMID: 33184579 Free PMC article.
-
Organoids as a Model for Colorectal Cancer.Curr Colorectal Cancer Rep. 2016;12(5):281-287. doi: 10.1007/s11888-016-0335-4. Epub 2016 Aug 6. Curr Colorectal Cancer Rep. 2016. PMID: 27656116 Free PMC article. Review.
-
Cell-type-specific signaling networks in heterocellular organoids.Nat Methods. 2020 Mar;17(3):335-342. doi: 10.1038/s41592-020-0737-8. Epub 2020 Feb 17. Nat Methods. 2020. PMID: 32066960 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

