Revealing Different Roles of the mTOR-Targets S6K1 and S6K2 in Breast Cancer by Expression Profiling and Structural Analysis
- PMID: 26698305
- PMCID: PMC4689523
- DOI: 10.1371/journal.pone.0145013
Revealing Different Roles of the mTOR-Targets S6K1 and S6K2 in Breast Cancer by Expression Profiling and Structural Analysis
Abstract
Background: The AKT/mTORC1/S6K pathway is frequently overstimulated in breast cancer, constituting a promising therapeutic target. The benefit from mTOR inhibitors varies, likely as a consequence of tumour heterogeneity, and upregulation of several compensatory feed-back mechanisms. The mTORC1 downstream effectors S6K1, S6K2, and 4EBP1 are amplified and overexpressed in breast cancer, associated with a poor outcome and divergent endocrine treatment benefit. S6K1 and S6K2 share high sequence homology, but evidence of partly distinct biological functions is emerging. The aim of this work was to explore possible different roles and treatment target potentials of S6K1 and S6K2 in breast cancer.
Materials and methods: Whole-genome expression profiles were compared for breast tumours expressing high levels of S6K1, S6K2 or 4EBP1, using public datasets, as well as after in vitro siRNA downregulation of S6K1 and/or S6K2 in ZR751 breast cancer cells. In silico homology modelling of the S6K2 kinase domain was used to evaluate its possible structural divergences to S6K1.
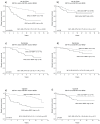
Results: Genome expression profiles were highly different in S6K1 and S6K2 high tumours, whereas S6K2 and 4EBP1 profiles showed significant overlaps, both correlated to genes involved in cell cycle progression, among these the master regulator E2F1. S6K2 and 4EBP1 were inversely associated with IGF1 levels, and their prognostic value was shown to be restricted to tumours positive for IGFR and/or HER2. In vitro, S6K1 and S6K2 silencing resulted in upregulation of genes in the mTORC1 and mTORC2 complexes. Isoform-specific silencing also showed distinct patterns, e.g. S6K2 downregulation lead to upregulation of several cell cycle associated genes. Structural analyses of the S6K2 kinase domain showed unique structure patterns, deviating from those of S6K1, facilitating the development of isoform-specific inhibitors. Our data support emerging proposals of distinct biological features of S6K1 and S6K2, suggesting their importance as separate oncogenes and clinical markers, where specific targeting in different breast cancer subtypes could facilitate further individualised therapies.
Conflict of interest statement
Figures


Similar articles
-
Distinct Roles of mTOR Targets S6K1 and S6K2 in Breast Cancer.Int J Mol Sci. 2020 Feb 11;21(4):1199. doi: 10.3390/ijms21041199. Int J Mol Sci. 2020. PMID: 32054043 Free PMC article. Review.
-
The mTOR effectors 4EBP1 and S6K2 are frequently coexpressed, and associated with a poor prognosis and endocrine resistance in breast cancer: a retrospective study including patients from the randomised Stockholm tamoxifen trials.Breast Cancer Res. 2013;15(5):R96. doi: 10.1186/bcr3557. Breast Cancer Res. 2013. PMID: 24131622 Free PMC article.
-
Clinical potential of the mTOR targets S6K1 and S6K2 in breast cancer.Breast Cancer Res Treat. 2011 Aug;128(3):713-23. doi: 10.1007/s10549-010-1058-x. Epub 2010 Oct 16. Breast Cancer Res Treat. 2011. PMID: 20953835
-
High-resolution genomic analysis of the 11q13 amplicon in breast cancers identifies synergy with 8p12 amplification, involving the mTOR targets S6K2 and 4EBP1.Genes Chromosomes Cancer. 2011 Oct;50(10):775-87. doi: 10.1002/gcc.20900. Epub 2011 Jul 11. Genes Chromosomes Cancer. 2011. PMID: 21748818
-
S6K2 promises an important therapeutic potential for cancer.Future Oncol. 2019 Jan;15(1):95-102. doi: 10.2217/fon-2018-0332. Epub 2018 Nov 1. Future Oncol. 2019. PMID: 30730779 Review.
Cited by
-
Transcriptomic analysis reveals the oncogenic role of S6K1 in hepatocellular carcinoma.J Cancer. 2020 Feb 19;11(9):2645-2655. doi: 10.7150/jca.40726. eCollection 2020. J Cancer. 2020. PMID: 32201535 Free PMC article.
-
mTORC1 induces eukaryotic translation initiation factor 4E interaction with TOS-S6 kinase 1 and its activation.Cell Cycle. 2021 May;20(9):839-854. doi: 10.1080/15384101.2021.1901038. Epub 2021 May 3. Cell Cycle. 2021. PMID: 33938392 Free PMC article.
-
Leptin Regulation of Cancer Stem Cells in Breast and Gynecologic Cancer.Endocrinology. 2018 Aug 1;159(8):3069-3080. doi: 10.1210/en.2018-00379. Endocrinology. 2018. PMID: 29955847 Free PMC article. Review.
-
mEAK-7 Forms an Alternative mTOR Complex with DNA-PKcs in Human Cancer.iScience. 2019 Jul 26;17:190-207. doi: 10.1016/j.isci.2019.06.029. Epub 2019 Jun 25. iScience. 2019. PMID: 31288154 Free PMC article.
-
Distinct Roles of mTOR Targets S6K1 and S6K2 in Breast Cancer.Int J Mol Sci. 2020 Feb 11;21(4):1199. doi: 10.3390/ijms21041199. Int J Mol Sci. 2020. PMID: 32054043 Free PMC article. Review.
References
-
- Sarbassov DD, Guertin DA, Ali SM, Sabatini DM (2005) Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307: 1098–1101. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

