Distinctive Behaviors of Druggable Proteins in Cellular Networks
- PMID: 26699810
- PMCID: PMC4689399
- DOI: 10.1371/journal.pcbi.1004597
Distinctive Behaviors of Druggable Proteins in Cellular Networks
Abstract
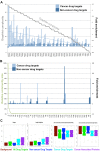
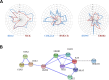
The interaction environment of a protein in a cellular network is important in defining the role that the protein plays in the system as a whole, and thus its potential suitability as a drug target. Despite the importance of the network environment, it is neglected during target selection for drug discovery. Here, we present the first systematic, comprehensive computational analysis of topological, community and graphical network parameters of the human interactome and identify discriminatory network patterns that strongly distinguish drug targets from the interactome as a whole. Importantly, we identify striking differences in the network behavior of targets of cancer drugs versus targets from other therapeutic areas and explore how they may relate to successful drug combinations to overcome acquired resistance to cancer drugs. We develop, computationally validate and provide the first public domain predictive algorithm for identifying druggable neighborhoods based on network parameters. We also make available full predictions for 13,345 proteins to aid target selection for drug discovery. All target predictions are available through canSAR.icr.ac.uk. Underlying data and tools are available at https://cansar.icr.ac.uk/cansar/publications/druggable_network_neighbourhoods/.
Conflict of interest statement
I have read the journal's policy and the authors of this manuscript have the following competing interests: The authors work at The Institute of Cancer Research, London, UK, which has a commercial interest in the discovery and development of anticancer drugs.
Figures

Similar articles
-
canSAR: an updated cancer research and drug discovery knowledgebase.Nucleic Acids Res. 2016 Jan 4;44(D1):D938-43. doi: 10.1093/nar/gkv1030. Epub 2015 Dec 15. Nucleic Acids Res. 2016. PMID: 26673713 Free PMC article.
-
canSAR: update to the cancer translational research and drug discovery knowledgebase.Nucleic Acids Res. 2019 Jan 8;47(D1):D917-D922. doi: 10.1093/nar/gky1129. Nucleic Acids Res. 2019. PMID: 30496479 Free PMC article.
-
Successful anti-cancer drug targets able to pass FDA review demonstrate the identifiable signature distinct from the signatures of random genes and initially proposed targets.Bioinformatics. 2008 Feb 1;24(3):389-95. doi: 10.1093/bioinformatics/btm447. Epub 2007 Oct 8. Bioinformatics. 2008. PMID: 17925305
-
Strategies in functional proteomics: Unveiling the pathways to precision oncology.Cancer Lett. 2016 Nov 1;382(1):86-94. doi: 10.1016/j.canlet.2016.01.049. Epub 2016 Feb 2. Cancer Lett. 2016. PMID: 26850375 Review.
-
Inside the biochemical pathways of thymidylate synthase perturbed by anticancer drugs: Novel strategies to overcome cancer chemoresistance.Drug Resist Updat. 2015 Nov;23:20-54. doi: 10.1016/j.drup.2015.10.003. Epub 2015 Oct 31. Drug Resist Updat. 2015. PMID: 26690339 Review.
Cited by
-
Screening out irrelevant cell-based models of disease.Nat Rev Drug Discov. 2016 Nov;15(11):751-769. doi: 10.1038/nrd.2016.175. Epub 2016 Sep 12. Nat Rev Drug Discov. 2016. PMID: 27616293 Review.
-
Whole-genome sequencing of multiple myeloma reveals oncogenic pathways are targeted somatically through multiple mechanisms.Leukemia. 2018 Nov;32(11):2459-2470. doi: 10.1038/s41375-018-0103-3. Epub 2018 Apr 9. Leukemia. 2018. PMID: 29654271 Free PMC article.
-
Biomarker-Drug and Liquid Biopsy Co-development for Disease Staging and Targeted Therapy: Cornerstones for Alzheimer's Precision Medicine and Pharmacology.Front Pharmacol. 2019 Mar 29;10:310. doi: 10.3389/fphar.2019.00310. eCollection 2019. Front Pharmacol. 2019. PMID: 30984002 Free PMC article.
-
Systematic interrogation of diverse Omic data reveals interpretable, robust, and generalizable transcriptomic features of clinically successful therapeutic targets.PLoS Comput Biol. 2018 May 21;14(5):e1006142. doi: 10.1371/journal.pcbi.1006142. eCollection 2018 May. PLoS Comput Biol. 2018. PMID: 29782487 Free PMC article.
-
MEDICI: Mining Essentiality Data to Identify Critical Interactions for Cancer Drug Target Discovery and Development.PLoS One. 2017 Jan 24;12(1):e0170339. doi: 10.1371/journal.pone.0170339. eCollection 2017. PLoS One. 2017. PMID: 28118365 Free PMC article.
References
-
- Hopkins A, Groom C (2002) The druggable genome. Nat Rev Drug Discov 1(9): 727–730. - PubMed
-
- Hajduk PJ, Huth JR, Fesik SW (2005) Druggability indices for protein targets derived from NMR-based screening data. J Med Chem 48: 2518–2525. - PubMed
-
- Al-Lazikani B, Gaulton A, Paolini GV, Lanfear J, Overington JP, et al. (2007) The Holy Grail: Molecular Function—The Molecular Basis of Predicting Druggability Bioinformatics: From Genomes to Therapies. Wiley-VCH, Vol. 3.
-
- Keller TH, Pichota A, Yin Z (2006) A practical view of 'druggability'. Curr Opin Chem Biol 10(4): 357–361. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

