Defective control of pre-messenger RNA splicing in human disease
- PMID: 26728853
- PMCID: PMC4700483
- DOI: 10.1083/jcb.201510032
Defective control of pre-messenger RNA splicing in human disease
Abstract
Examples of associations between human disease and defects in pre-messenger RNA splicing/alternative splicing are accumulating. Although many alterations are caused by mutations in splicing signals or regulatory sequence elements, recent studies have noted the disruptive impact of mutated generic spliceosome components and splicing regulatory proteins. This review highlights recent progress in our understanding of how the altered splicing function of RNA-binding proteins contributes to myelodysplastic syndromes, cancer, and neuropathologies.
© 2016 Chabot and Shkreta.
Figures
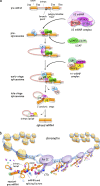
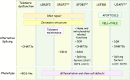

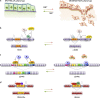


References
-
- Adler A.S., McCleland M.L., Yee S., Yaylaoglu M., Hussain S., Cosino E., Quinones G., Modrusan Z., Seshagiri S., Torres E., et al. 2014. An integrative analysis of colon cancer identifies an essential function for PRPF6 in tumor growth. Genes Dev. 28:1068–1084. 10.1101/gad.237206.113 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

