Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB and insights into the mechanism of catalysis
- PMID: 26729507
- PMCID: PMC4728340
- DOI: 10.1038/ncomms10175
Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB and insights into the mechanism of catalysis
Abstract
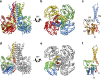
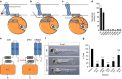
The attachment of a sugar to a hydrophobic polyisoprenyl carrier is the first step for all extracellular glycosylation processes. The enzymes that perform these reactions, polyisoprenyl-glycosyltransferases (PI-GTs) include dolichol phosphate mannose synthase (DPMS), which generates the mannose donor for glycosylation in the endoplasmic reticulum. Here we report the 3.0 Å resolution crystal structure of GtrB, a glucose-specific PI-GT from Synechocystis, showing a tetramer in which each protomer contributes two helices to a membrane-spanning bundle. The active site is 15 Å from the membrane, raising the question of how water-soluble and membrane-embedded substrates are brought into apposition for catalysis. A conserved juxtamembrane domain harbours disease mutations, which compromised activity in GtrB in vitro and in human DPM1 tested in zebrafish. We hypothesize a role of this domain in shielding the polyisoprenyl-phosphate for transport to the active site. Our results reveal the basis of PI-GT function, and provide a potential molecular explanation for DPM1-related disease.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


Similar articles
-
Dolichol phosphate mannose synthase: a Glycosyltransferase with Unity in molecular diversities.Glycoconj J. 2017 Aug;34(4):467-479. doi: 10.1007/s10719-017-9777-4. Epub 2017 Jun 14. Glycoconj J. 2017. PMID: 28616799 Free PMC article. Review.
-
Enzymes that recognize dolichols participate in three glycosylation pathways and are required for protein secretion.Biochem Cell Biol. 1992 Jun;70(6):438-47. doi: 10.1139/o92-067. Biochem Cell Biol. 1992. PMID: 1333231 Review.
-
Structure and mechanism of the ER-based glucosyltransferase ALG6.Nature. 2020 Mar;579(7799):443-447. doi: 10.1038/s41586-020-2044-z. Epub 2020 Feb 26. Nature. 2020. PMID: 32103179 Free PMC article.
-
A Genome-Wide CRISPR-Cas9 Screen Identifies the Dolichol-Phosphate Mannose Synthase Complex as a Host Dependency Factor for Dengue Virus Infection.J Virol. 2020 Mar 17;94(7):e01751-19. doi: 10.1128/JVI.01751-19. Print 2020 Mar 17. J Virol. 2020. PMID: 31915280 Free PMC article.
-
Distinct Regions of the Haloferax volcanii Dolichol Phosphate-Mannose Synthase AglD Mediate the Assembly and Subsequent Processing of the Lipid-Linked Mannose.J Bacteriol. 2022 Jan 18;204(1):e0044721. doi: 10.1128/JB.00447-21. Epub 2021 Oct 11. J Bacteriol. 2022. PMID: 34633871 Free PMC article.
Cited by
-
Molecular mechanisms underlying the structural diversity of rhamnose-rich cell wall polysaccharides in lactococci.J Biol Chem. 2024 Jan;300(1):105578. doi: 10.1016/j.jbc.2023.105578. Epub 2023 Dec 16. J Biol Chem. 2024. PMID: 38110036 Free PMC article.
-
Structural basis of undecaprenyl phosphate glycosylation leading to polymyxin resistance in Gram-negative bacteria.bioRxiv [Preprint]. 2025 Jan 30:2025.01.29.634835. doi: 10.1101/2025.01.29.634835. bioRxiv. 2025. PMID: 39974898 Free PMC article. Preprint.
-
Making the Enterobacterial Common Antigen Glycan and Measuring Its Substrate Sequestration.ACS Chem Biol. 2021 Apr 16;16(4):691-700. doi: 10.1021/acschembio.0c00983. Epub 2021 Mar 19. ACS Chem Biol. 2021. PMID: 33740380 Free PMC article.
-
Dolichol phosphate mannose synthase: a Glycosyltransferase with Unity in molecular diversities.Glycoconj J. 2017 Aug;34(4):467-479. doi: 10.1007/s10719-017-9777-4. Epub 2017 Jun 14. Glycoconj J. 2017. PMID: 28616799 Free PMC article. Review.
-
Molecular Cloning, Expression and Characterization of Oenococcus oeni Priming Glycosyltransferases.Mol Biotechnol. 2017 Aug;59(8):323-333. doi: 10.1007/s12033-017-0021-z. Mol Biotechnol. 2017. PMID: 28667570
References
-
- Allison G. E. & Verma N. K. Serotype-converting bacteriophages and O-antigen modification in Shigella flexneri. Trends Microbiol. 8, 17–23 (2000). - PubMed
-
- Bugg T. D. & Brandish P. E. From peptidoglycan to glycoproteins: common features of lipid-linked oligosaccharide biosynthesis. FEMS Microbiol. Lett. 119, 255–262 (1994). - PubMed
-
- Jensen J. W. & Schutzbach J. S. Characterization of mannosyl-transfer reactions catalyzed by dolichyl-mannosyl-phosphate-synthase. Carbohydr. Res. 149, 199–208 (1986). - PubMed
-
- Maeda Y. & Kinoshita T. Dolichol-phosphate mannose synthase: Structure, function and regulation. Biochim. Biophys. Acta 1780, 861–868 (2008). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

