Origins and Evolution of Hepatitis B Virus and Hepatitis D Virus
- PMID: 26729756
- PMCID: PMC4691804
- DOI: 10.1101/cshperspect.a021360
Origins and Evolution of Hepatitis B Virus and Hepatitis D Virus
Abstract
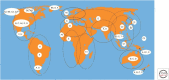
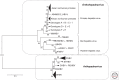
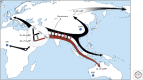
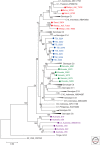
Members of the family Hepadnaviridae fall into two subgroups: mammalian and avian. The detection of endogenous avian hepadnavirus DNA integrated into the genomes of zebra finches has revealed a deep evolutionary origin of hepadnaviruses that was not previously recognized, dating back at least 40 million and possibly >80 million years ago. The nonprimate mammalian members of the Hepadnaviridae include the woodchuck hepatitis virus (WHV), the ground squirrel hepatitis virus, and arctic squirrel hepatitis virus, as well as a number of members of the recently described bat hepatitis virus. The identification of hepatitis B viruses (HBVs) in higher primates, such as chimpanzee, gorilla, orangutan, and gibbons that cluster with the human HBV, as well as a number of recombinant forms between humans and primates, further implies a more complex origin of this virus. We discuss the current theories of the origin and evolution of HBV and propose a model that includes cross-species transmissions and subsequent recombination events on a genetic backbone of genotype C HBV infection. The hepatitis delta virus (HDV) is a defective RNA virus requiring the presence of the HBV for the completion of its life cycle. The origins of this virus remain unknown, although some recent studies have suggested an ancient African radiation. The age of the association between HDV and HBV is also unknown.
Copyright © 2016 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures


References
-
- Acharyya S, Basu P. 1993. Toba ash on the Indian subcontinent and its implications for correlation of late pleistocene alluvium. Quat Int 40: 10–19.
-
- Ahn SH, Yuen L, Han KH, Littlejohn M, Chang HY, Damerow H, Ayres A, Heo J, Locarnini S, Revill PA. 2010. Molecular and clinical characteristics of hepatitis B virus in Korea. J Med Virol 82: 1126–1134. - PubMed
-
- Banerjee A, Kurbanov F, Datta S, Chandra PK, Tanaka Y, Mizokami M, Chakravarty R. 2006. Phylogenetic relatedness and genetic diversity of hepatitis B virus isolates in Eastern India. J Med Virol 78: 1164–1174. - PubMed
-
- Bollyky P, Rambaut A, Grassly NC, et al. 1997. Hepatitis B virus has a New World evolutionary origin. Hepatology 26: 765.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
