Quantitative Phosphoproteomics Analysis of ERBB3/ERBB4 Signaling
- PMID: 26745281
- PMCID: PMC4706443
- DOI: 10.1371/journal.pone.0146100
Quantitative Phosphoproteomics Analysis of ERBB3/ERBB4 Signaling
Abstract
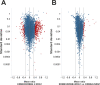
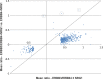
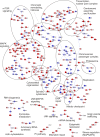
The four members of the epidermal growth factor receptor (EGFR/ERBB) family form homo- and heterodimers which mediate ligand-specific regulation of many key cellular processes in normal and cancer tissues. While signaling through the EGFR has been extensively studied on the molecular level, signal transduction through ERBB3/ERBB4 heterodimers is less well understood. Here, we generated isogenic mouse Ba/F3 cells that express full-length and functional membrane-integrated ERBB3 and ERBB4 or ERBB4 alone, to serve as a defined cellular model for biological and phosphoproteomics analysis of ERBB3/ERBB4 signaling. ERBB3 co-expression significantly enhanced Ba/F3 cell proliferation upon neuregulin-1 (NRG1) treatment. For comprehensive signaling studies we performed quantitative mass spectrometry (MS) experiments to compare the basal ERBB3/ERBB4 cell phosphoproteome to NRG1 treatment of ERBB3/ERBB4 and ERBB4 cells. We employed a workflow comprising differential isotope labeling with mTRAQ reagents followed by chromatographic peptide separation and final phosphopeptide enrichment prior to MS analysis. Overall, we identified 9686 phosphorylation sites which could be confidently localized to specific residues. Statistical analysis of three replicate experiments revealed 492 phosphorylation sites which were significantly changed in NRG1-treated ERBB3/ERBB4 cells. Bioinformatics data analysis recapitulated regulation of mitogen-activated protein kinase and Akt pathways, but also indicated signaling links to cytoskeletal functions and nuclear biology. Comparative assessment of NRG1-stimulated ERBB4 Ba/F3 cells revealed that ERBB3 did not trigger defined signaling pathways but more broadly enhanced phosphoproteome regulation in cells expressing both receptors. In conclusion, our data provide the first global picture of ERBB3/ERBB4 signaling and provide numerous potential starting points for further mechanistic studies.
Conflict of interest statement
Figures


Similar articles
-
Neuregulin-1 only induces trans-phosphorylation between ErbB receptor heterodimer partners.Cell Signal. 2007 Mar;19(3):466-71. doi: 10.1016/j.cellsig.2006.07.020. Epub 2006 Sep 15. Cell Signal. 2007. PMID: 16978839
-
An ERBB3/ERBB2 oncogenic unit plays a key role in NRG1 signaling and melanoma cell growth and survival.Pigment Cell Melanoma Res. 2013 May;26(3):408-14. doi: 10.1111/pcmr.12089. Epub 2013 Apr 2. Pigment Cell Melanoma Res. 2013. PMID: 23480537
-
ErbB2 and ErbB3 do not quantitatively modulate ligand-induced ErbB4 tyrosine phosphorylation.Cell Signal. 2002 Sep;14(9):793-8. doi: 10.1016/s0898-6568(02)00019-0. Cell Signal. 2002. PMID: 12034361
-
The neuregulin-I/ErbB signaling system in development and disease.Adv Anat Embryol Cell Biol. 2007;190:1-65. Adv Anat Embryol Cell Biol. 2007. PMID: 17432114 Review.
-
Neuregulin-1 and ALS19 (ERBB4): at the crossroads of amyotrophic lateral sclerosis and cancer.BMC Med. 2024 Feb 19;22(1):74. doi: 10.1186/s12916-024-03293-3. BMC Med. 2024. PMID: 38369520 Free PMC article. Review.
Cited by
-
Activated Protein Kinase C (PKC) Is Persistently Trafficked with Epidermal Growth Factor (EGF) Receptor.Biomolecules. 2020 Sep 7;10(9):1288. doi: 10.3390/biom10091288. Biomolecules. 2020. PMID: 32906765 Free PMC article.
-
Community detection in empirical kinase networks identifies new potential members of signalling pathways.PLoS Comput Biol. 2023 Jun 23;19(6):e1010459. doi: 10.1371/journal.pcbi.1010459. eCollection 2023 Jun. PLoS Comput Biol. 2023. PMID: 37352361 Free PMC article.
-
Investigation of Rare Non-Coding Variants in Familial Multiple Myeloma.Cells. 2022 Dec 26;12(1):96. doi: 10.3390/cells12010096. Cells. 2022. PMID: 36611892 Free PMC article.
-
Treadmill Running Regulates Adult Neurogenesis, Spatial and Non-spatial Learning, Parvalbumin Neuron Activity by ErbB4 Signaling.Cell Mol Neurobiol. 2024 Jan 29;44(1):17. doi: 10.1007/s10571-023-01439-0. Cell Mol Neurobiol. 2024. PMID: 38285192 Free PMC article.
-
The silent spread: exploring diverse metastatic pathways in high-grade serous ovarian cancer.Front Med (Lausanne). 2025 Mar 5;12:1539024. doi: 10.3389/fmed.2025.1539024. eCollection 2025. Front Med (Lausanne). 2025. PMID: 40109727 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

