Introgression of Neandertal- and Denisovan-like Haplotypes Contributes to Adaptive Variation in Human Toll-like Receptors
- PMID: 26748514
- PMCID: PMC4716682
- DOI: 10.1016/j.ajhg.2015.11.015
Introgression of Neandertal- and Denisovan-like Haplotypes Contributes to Adaptive Variation in Human Toll-like Receptors
Erratum in
- Am J Hum Genet. 2016 Feb 4;98(2):399
-
Erratum: Introgression of Neandertal- and Denisovan-like Haplotypes Contributes to Adaptive Variation in Human Toll-like Receptors.Am J Hum Genet. 2016 Feb 4;98(2):399. doi: 10.1016/j.ajhg.2016.01.012. Am J Hum Genet. 2016. PMID: 31265485 Free PMC article.
Abstract
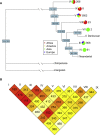
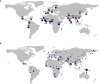
Pathogens and the diseases they cause have been among the most important selective forces experienced by humans during their evolutionary history. Although adaptive alleles generally arise by mutation, introgression can also be a valuable source of beneficial alleles. Archaic humans, who lived in Europe and Western Asia for more than 200,000 years, were probably well adapted to this environment and its local pathogens. It is therefore conceivable that modern humans entering Europe and Western Asia who admixed with them obtained a substantial immune advantage from the introgression of archaic alleles. Here we document a cluster of three Toll-like receptors (TLR6-TLR1-TLR10) in modern humans that carries three distinct archaic haplotypes, indicating repeated introgression from archaic humans. Two of these haplotypes are most similar to the Neandertal genome, and the third haplotype is most similar to the Denisovan genome. The Toll-like receptors are key components of innate immunity and provide an important first line of immune defense against bacteria, fungi, and parasites. The unusually high allele frequencies and unexpected levels of population differentiation indicate that there has been local positive selection on multiple haplotypes at this locus. We show that the introgressed alleles have clear functional effects in modern humans; archaic-like alleles underlie differences in the expression of the TLR genes and are associated with increased [corrected] microbial resistance and increased allergic disease in large cohorts. This provides strong evidence for recurrent adaptive introgression at the TLR6-TLR1-TLR10 locus, resulting in differences in disease phenotypes in modern humans.
Copyright © 2016 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



Comment in
-
TLRs of Our Fathers.Immunity. 2016 Feb 16;44(2):218-20. doi: 10.1016/j.immuni.2016.02.003. Immunity. 2016. PMID: 26885854
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

