Spontaneous cleavage of proteins at serine and threonine is facilitated by zinc
- PMID: 26751411
- PMCID: PMC4783340
- DOI: 10.1111/acel.12428
Spontaneous cleavage of proteins at serine and threonine is facilitated by zinc
Abstract
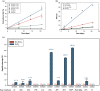
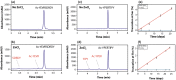
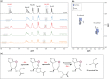
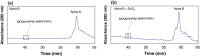
Old proteins are widely distributed in the body. Over time, they deteriorate and many spontaneous reactions, for example isomerisation of Asp and Asn, can be replicated by incubation of peptides under physiological conditions. One of the signatures of long-lived proteins that has proven to be difficult to replicate in vitro is cleavage on the N-terminal side of Ser residues, and this is important since cleavage at Ser, and also Thr, has been observed in a number of human proteins. In this study, the autolysis of Ser- and Thr-containing peptides was investigated with particular reference to discovering factors that promote cleavage adjacent to Ser/Thr at neutral pH. It was found that zinc catalyses cleavage of the peptide bond on the N-terminal side of Ser residues and further that this process is markedly accelerated if a His residue is adjacent to the Ser. NMR analysis indicated that the imidazole group co-ordinates zinc and that once zinc is co-ordinated, it can polarize the carbonyl group of the peptide bond in a manner analogous to that observed in the active site of the metalloexopeptidase, carboxypeptidase A. The hydroxyl side chain of Ser/Thr is then able to cleave the adjacent peptide bond. These observations enable an understanding of the origin of common truncations observed in long-lived proteins, for example truncation on the N-terminal side of Ser 8 in Abeta, Ser 19 in alpha B crystallin and Ser 66 in alpha A crystallin. The presence of zinc may therefore significantly affect the long-term stability of cellular proteins.
Keywords: Alzheimer's disease; ageing; aging; lifespan; longevity; neurodegenerative disease; protein chemistry; proteomics.
© 2016 The Authors. Aging Cell published by the Anatomical Society and John Wiley & Sons Ltd.
Figures


Similar articles
-
Cleavage C-terminal to Asp leads to covalent crosslinking of long-lived human proteins.Biochim Biophys Acta Proteins Proteom. 2019 Sep;1867(9):831-839. doi: 10.1016/j.bbapap.2019.06.009. Epub 2019 Jun 19. Biochim Biophys Acta Proteins Proteom. 2019. PMID: 31226490 Free PMC article.
-
Analysis of protein phosphorylation in the regions of consecutive serine/threonine residues by negative ion electrospray collision-induced dissociation. Approach to pinpointing of phosphorylation sites.Anal Chem. 2007 May 1;79(9):3476-86. doi: 10.1021/ac0623991. Epub 2007 Mar 28. Anal Chem. 2007. PMID: 17388569
-
Hydrogen exchange study on the hydroxyl groups of serine and threonine residues in proteins and structure refinement using NOE restraints with polar side-chain groups.J Am Chem Soc. 2011 Nov 2;133(43):17420-7. doi: 10.1021/ja206799v. Epub 2011 Oct 10. J Am Chem Soc. 2011. PMID: 21955241
-
Collision-induced fragmentations of the (M-H)- parent anions of underivatized peptides: an aid to structure determination and some unusual negative ion cleavages.Mass Spectrom Rev. 2002 Mar-Apr;21(2):87-107. doi: 10.1002/mas.10022. Mass Spectrom Rev. 2002. PMID: 12373746 Review.
-
Serine/threonine ligation for the chemical synthesis of proteins.Curr Opin Chem Biol. 2014 Oct;22:108-14. doi: 10.1016/j.cbpa.2014.09.023. Epub 2014 Oct 6. Curr Opin Chem Biol. 2014. PMID: 25299572 Review.
Cited by
-
Identification and Functional Assessment of Age-Dependent Truncations to Cx46 and Cx50 in the Human Lens.Invest Ophthalmol Vis Sci. 2016 Oct 1;57(13):5714-5722. doi: 10.1167/iovs.16-19698. Invest Ophthalmol Vis Sci. 2016. PMID: 27787559 Free PMC article.
-
Spontaneous Cleavage at Glu and Gln Residues in Long-Lived Proteins.ACS Chem Biol. 2021 Nov 19;16(11):2244-2254. doi: 10.1021/acschembio.1c00379. Epub 2021 Oct 22. ACS Chem Biol. 2021. PMID: 34677941 Free PMC article.
-
Molecular Processes Implicated in Human Age-Related Nuclear Cataract.Invest Ophthalmol Vis Sci. 2019 Dec 2;60(15):5007-5021. doi: 10.1167/iovs.19-27535. Invest Ophthalmol Vis Sci. 2019. PMID: 31791064 Free PMC article. Review.
-
Quantification of serine residue stereoinversion in a short peptide by reversed-phase high-performance liquid chromatography: analysis of mechanisms promoting serine stereoinversion.Anal Sci. 2024 May;40(5):925-934. doi: 10.1007/s44211-024-00543-5. Epub 2024 Mar 26. Anal Sci. 2024. PMID: 38528254
-
Can the Fact That Myelin Proteins Are Old and Break down Explain the Origin of Multiple Sclerosis in Some People?J Clin Med. 2018 Sep 14;7(9):281. doi: 10.3390/jcm7090281. J Clin Med. 2018. PMID: 30223497 Free PMC article. Review.
References
-
- Cloos PA, Christgau S (2002) Non‐enzymatic covalent modifications of proteins: mechanisms, physiological consequences and clinical applications. Matrix Biol. 21, 39–52. - PubMed
-
- Cohen SL, Price C, Vlasak J (2007) β‐Elimination and peptide bond hydrolysis: two distinct mechanisms of human IgG1 hinge fragmentation upon storage. J. Am. Chem. Soc. 129, 6976–6977. - PubMed
-
- Duncan EA, Brown MS, Goldstein JL, Sakai J (1997) Cleavage site for sterol‐regulated protease localized to a leu‐Ser bond in the lumenal loop of sterol regulatory element‐binding protein‐2. J Biol Chem. 272, 12778‐12785. http://www.ncbi.nlm.nih.gov/pubmed/9139737 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

