GBS-SNP-CROP: a reference-optional pipeline for SNP discovery and plant germplasm characterization using variable length, paired-end genotyping-by-sequencing data
- PMID: 26754002
- PMCID: PMC4709900
- DOI: 10.1186/s12859-016-0879-y
GBS-SNP-CROP: a reference-optional pipeline for SNP discovery and plant germplasm characterization using variable length, paired-end genotyping-by-sequencing data
Abstract
Background: With its simple library preparation and robust approach to genome reduction, genotyping-by-sequencing (GBS) is a flexible and cost-effective strategy for SNP discovery and genotyping, provided an appropriate reference genome is available. For resource-limited curation, research, and breeding programs of underutilized plant genetic resources, however, even low-depth references may not be within reach, despite declining sequencing costs. Such programs would find value in an open-source bioinformatics pipeline that can maximize GBS data usage and perform high-density SNP genotyping in the absence of a reference.
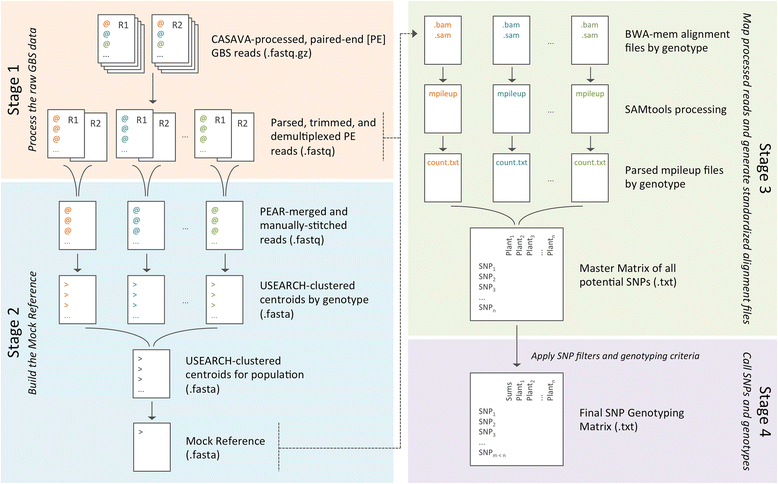
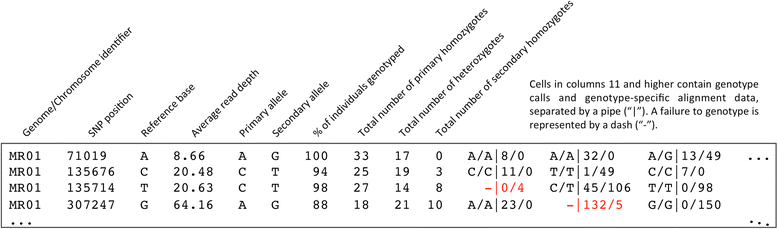
Results: The GBS SNP-Calling Reference Optional Pipeline (GBS-SNP-CROP) developed and presented here adopts a clustering strategy to build a population-tailored "Mock Reference" from the same GBS data used for downstream SNP calling and genotyping. Designed for libraries of paired-end (PE) reads, GBS-SNP-CROP maximizes data usage by eliminating unnecessary data culling due to imposed read-length uniformity requirements. Using 150 bp PE reads from a GBS library of 48 accessions of tetraploid kiwiberry (Actinidia arguta), GBS-SNP-CROP yielded on average three times as many SNPs as TASSEL-GBS analyses (32 and 64 bp tag lengths) and over 18 times as many as TASSEL-UNEAK, with fewer genotyping errors in all cases, as evidenced by comparing the genotypic characterizations of biological replicates. Using the published reference genome of a related diploid species (A. chinensis), the reference-based version of GBS-SNP-CROP behaved similarly to TASSEL-GBS in terms of the number of SNPs called but had an improved read depth distribution and fewer genotyping errors. Our results also indicate that the sets of SNPs detected by the different pipelines above are largely orthogonal to one another; thus GBS-SNP-CROP may be used to augment the results of alternative analyses, whether or not a reference is available.
Conclusions: By achieving high-density SNP genotyping in populations for which no reference genome is available, GBS-SNP-CROP is worth consideration by curators, researchers, and breeders of under-researched plant genetic resources. In cases where a reference is available, especially if from a related species or when the target population is particularly diverse, GBS-SNP-CROP may complement other reference-based pipelines by extracting more information per sequencing dollar spent. The current version of GBS-SNP-CROP is available at https://github.com/halelab/GBS-SNP-CROP.git.
Figures

References
-
- Naylor RL, Falcona WP, Goodmanb RM, Jahnc MM, Sengoobad T, Teferae H, et al. Biotechnology in the developing world: a case for increased investments in orphan crops. Food Policy. 2004;29(1):15–44. doi: 10.1016/j.foodpol.2004.01.002. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

