Regulatory Characteristics of Vibrio vulnificus gbpA Gene Encoding a Mucin-binding Protein Essential for Pathogenesis
- PMID: 26755724
- PMCID: PMC4786714
- DOI: 10.1074/jbc.M115.685321
Regulatory Characteristics of Vibrio vulnificus gbpA Gene Encoding a Mucin-binding Protein Essential for Pathogenesis
Abstract
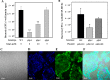
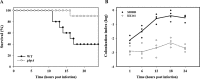
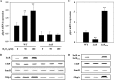
Binding to mucin is the initial step for enteropathogens to establish pathogenesis. An open reading frame, gbpA, of Vibrio vulnificus was identified and characterized in this study. Compared with wild type, the gbpA mutant was impaired in binding to mucin-agar and the mucin-secreting HT29-methotrexate cells, and the impaired mucin binding was restored by the purified GbpA provided exogenously. The gbpA mutant had attenuated virulence and ability of intestinal colonization in a mouse model, indicating that GbpA is a mucin-binding protein and essential for pathogenesis of V. vulnificus. The gbpA transcription was growth phase-dependent, reaching a maximum during the exponential phase. The Fe-S cluster regulator (IscR) and the cyclic AMP receptor protein (CRP) coactivated, whereas SmcR, a LuxR homologue, repressed gbpA. The cellular levels of IscR, CRP, and SmcR were not significantly affected by one another, indicating that the regulator proteins function cooperatively to regulate gbpA rather than sequentially in a regulatory cascade. The regulatory proteins directly bind upstream of the gbpA promoter PgbpA. DNase I protection assays, together with the deletion analyses of PgbpA, demonstrated that IscR binds to two specific sequences centered at -164.5 and -106, and CRP and SmcR bind specifically to the sequences centered at -68 and -45, respectively. Furthermore, gbpA was induced by exposure to H2O2, and the induction appeared to be mediated by elevated intracellular levels of IscR. Consequently, the combined results indicated that IscR, CRP, and SmcR cooperate for precise regulation of gbpA during the V. vulnificus pathogenesis.
Keywords: CRP; GbpA; IscR; LuxR homologue; Vibrio vulnificus; bacterial adhesion; gene regulation; mucin; oxidative stress; quorum sensing.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures







Similar articles
-
Characterization of the Vibrio vulnificus 1-Cys peroxiredoxin Prx3 and regulation of its expression by the Fe-S cluster regulator IscR in response to oxidative stress and iron starvation.J Biol Chem. 2014 Dec 26;289(52):36263-74. doi: 10.1074/jbc.M114.611020. Epub 2014 Nov 14. J Biol Chem. 2014. PMID: 25398878 Free PMC article.
-
LuxR homologue SmcR is essential for Vibrio vulnificus pathogenesis and biofilm detachment, and its expression is induced by host cells.Infect Immun. 2013 Oct;81(10):3721-30. doi: 10.1128/IAI.00561-13. Epub 2013 Jul 29. Infect Immun. 2013. PMID: 23897607 Free PMC article.
-
IscR is a global regulator essential for pathogenesis of Vibrio vulnificus and induced by host cells.Infect Immun. 2014 Feb;82(2):569-78. doi: 10.1128/IAI.01141-13. Epub 2013 Nov 18. Infect Immun. 2014. PMID: 24478072 Free PMC article.
-
Molecular Pathogenesis of Vibrio vulnificus.J Microbiol. 2005 Feb;43 Spec No:118-31. J Microbiol. 2005. PMID: 15765065 Review.
-
The regulator HlyU, the repeat-in-toxin gene rtxA1, and their roles in the pathogenesis of Vibrio vulnificus infections.Microbiologyopen. 2012 Dec;1(4):502-13. doi: 10.1002/mbo3.48. Epub 2012 Nov 28. Microbiologyopen. 2012. PMID: 23233275 Free PMC article. Review.
Cited by
-
Two novel genes identified by large-scale transcriptomic analysis are essential for biofilm and rugose colony development of Vibrio vulnificus.PLoS Pathog. 2023 Jan 19;19(1):e1011064. doi: 10.1371/journal.ppat.1011064. eCollection 2023 Jan. PLoS Pathog. 2023. PMID: 36656902 Free PMC article.
-
A Vibrio vulnificus VvpM Induces IL-1β Production Coupled with Necrotic Macrophage Death via Distinct Spatial Targeting by ANXA2.Front Cell Infect Microbiol. 2017 Aug 11;7:352. doi: 10.3389/fcimb.2017.00352. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28848713 Free PMC article.
-
Vibrio vulnificus RtxA Is a Major Factor Driving Inflammatory T Helper Type 17 Cell Responses in vitro and in vivo.Front Immunol. 2018 Sep 19;9:2095. doi: 10.3389/fimmu.2018.02095. eCollection 2018. Front Immunol. 2018. PMID: 30283443 Free PMC article.
-
Crystal Structure of the Regulatory Domain of AphB from Vibrio vulnificus, a Virulence Gene Regulator.Mol Cells. 2017 Apr;40(4):299-306. doi: 10.14348/molcells.2017.0015. Epub 2017 Apr 20. Mol Cells. 2017. PMID: 28427249 Free PMC article.
-
Effects of BlueM® against Streptococcus mutans biofilm and its virulence gene expression.Braz Dent J. 2023 Jan-Feb;34(1):19-28. doi: 10.1590/0103-6440202305133. Braz Dent J. 2023. PMID: 36888841 Free PMC article.
References
-
- McGuckin M. A., Lindén S. K., Sutton P., and Florin T. H. (2011) Mucin dynamics and enteric pathogens. Nat. Rev. Microbiol. 9, 265–278 - PubMed
-
- Neutra M. R., and Forstner J. F. (1987) in Physiology of the Gastrointestinal Tract (Johnson L. R., ed) Vol. 2, 2nd Ed., pp. 975–1009, Raven Press, Ltd., New York
-
- Thornton D. J., and Sheehan J. K. (2004) From mucins to mucus: toward a more coherent understanding of this essential barrier. Proc. Am. Thorac. Soc. 1, 54–61 - PubMed
-
- Ofek I., Bayer E. A., and Abraham S. N. (2013) in The Prokaryotes (Rosenberg E., Delong E. F., Lory S., Stackebrandt E., and Thompson F., eds) pp. 107–123, 10.1007/978-3-642-30144-5_50, Springer-Verlag, Berlin, Heidelberg - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

