Decision for cell fate: deubiquitinating enzymes in cell cycle checkpoint
- PMID: 26762302
- PMCID: PMC11108577
- DOI: 10.1007/s00018-015-2129-2
Decision for cell fate: deubiquitinating enzymes in cell cycle checkpoint
Abstract
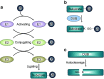
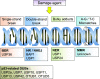
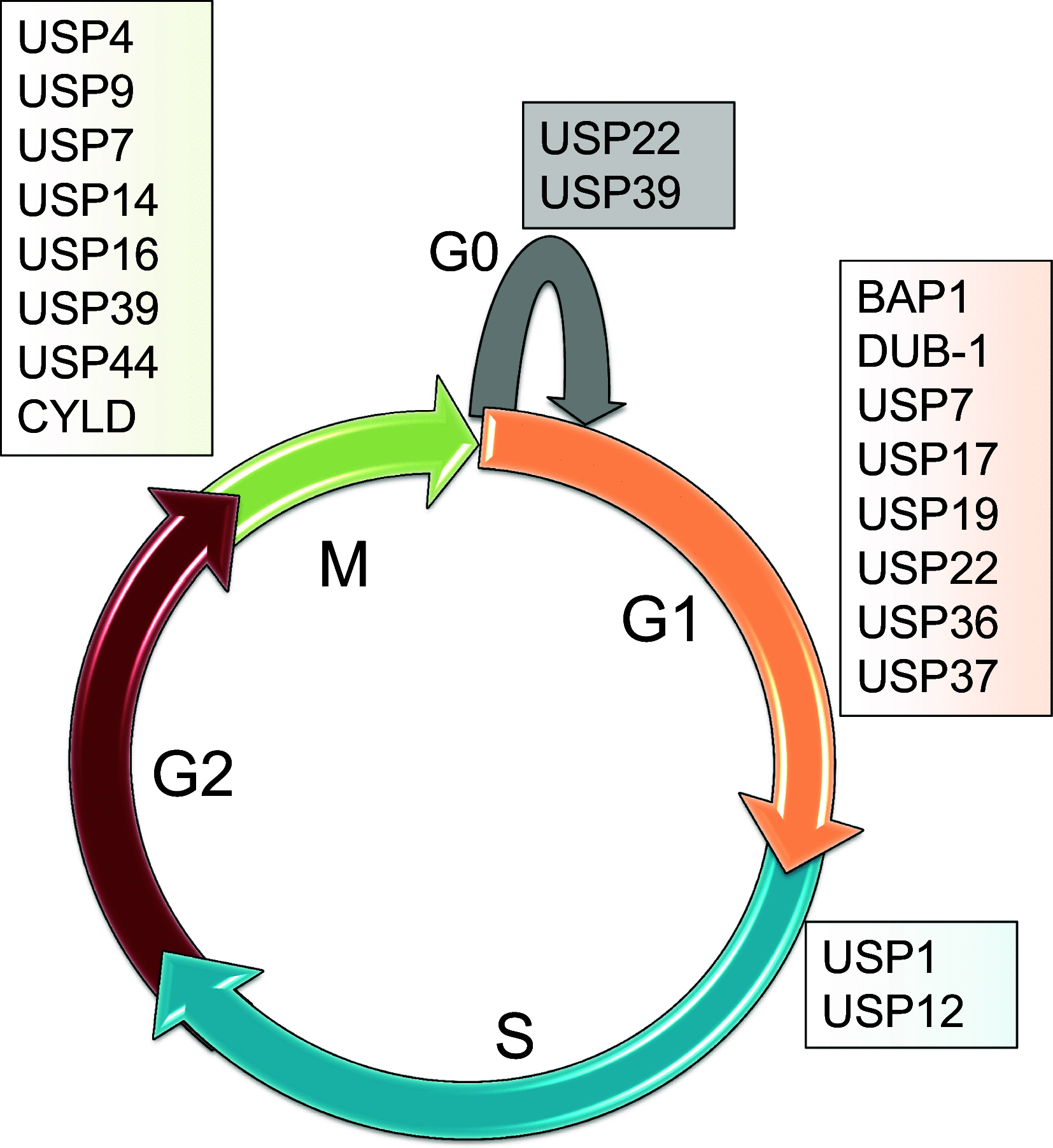
All organs consisting of single cells are consistently maintaining homeostasis in response to stimuli such as free oxygen, DNA damage, inflammation, and microorganisms. The cell cycle of all mammalian cells is regulated by protein expression in the right phase to respond to proliferation and apoptosis signals. Post-translational modifications (PTMs) of proteins by several protein-editing enzymes are associated with cell cycle regulation by their enzymatic functions. Ubiquitination, one of the PTMs, is also strongly related to cell cycle regulation by protein degradation or signal transduction. The importance of deubiquitinating enzymes (DUBs), which have a reversible function for ubiquitination, has recently suggested that the function of DUBs is also important for determining the fate of proteins during cell cycle processing. This article reviews and summarizes the diverse roles of DUBs, including DNA damage, cell cycle processing, and regulation of histone proteins, and also suggests the possibility for therapeutic targets.
Keywords: Cell cycle; DNA damage; Deubiquitinating enzyme; Ubiquitination.
Figures
Similar articles
-
Deubiquitinating enzymes as oncotargets.Oncotarget. 2015;6(12):9657-68. doi: 10.18632/oncotarget.3922. Oncotarget. 2015. PMID: 25962961 Free PMC article. Review.
-
H2A-DUBbing the mammalian epigenome: expanding frontiers for histone H2A deubiquitinating enzymes in cell biology and physiology.Int J Biochem Cell Biol. 2014 May;50:161-74. doi: 10.1016/j.biocel.2014.03.004. Epub 2014 Mar 16. Int J Biochem Cell Biol. 2014. PMID: 24647359 Review.
-
USP3 counteracts RNF168 via deubiquitinating H2A and γH2AX at lysine 13 and 15.Cell Cycle. 2014;13(1):106-14. doi: 10.4161/cc.26814. Epub 2013 Oct 24. Cell Cycle. 2014. PMID: 24196443 Free PMC article.
-
p53 stability is regulated by diverse deubiquitinating enzymes.Biochim Biophys Acta Rev Cancer. 2017 Dec;1868(2):404-411. doi: 10.1016/j.bbcan.2017.08.001. Epub 2017 Aug 8. Biochim Biophys Acta Rev Cancer. 2017. PMID: 28801249 Review.
-
Deubiquitylating enzymes and DNA damage response pathways.Cell Biochem Biophys. 2013 Sep;67(1):25-43. doi: 10.1007/s12013-013-9635-3. Cell Biochem Biophys. 2013. PMID: 23712866 Free PMC article. Review.
Cited by
-
Posttranslational control of lipogenesis in the tumor microenvironment.J Hematol Oncol. 2022 Aug 29;15(1):120. doi: 10.1186/s13045-022-01340-1. J Hematol Oncol. 2022. PMID: 36038892 Free PMC article. Review.
-
Identification of small-molecule binding sites of a ubiquitin-conjugating enzyme-UBE2T through fragment-based screening.Protein Sci. 2024 Mar;33(3):e4904. doi: 10.1002/pro.4904. Protein Sci. 2024. PMID: 38358126 Free PMC article.
-
DUBbing Cancer: Deubiquitylating Enzymes Involved in Epigenetics, DNA Damage and the Cell Cycle As Therapeutic Targets.Front Genet. 2016 Jul 28;7:133. doi: 10.3389/fgene.2016.00133. eCollection 2016. Front Genet. 2016. PMID: 27516771 Free PMC article. Review.
-
miR-543 impairs breast cancer cell phenotypes by targeting and suppressing ubiquitin-conjugating enzyme E2T (UBE2T).Bioengineered. 2021 Dec;12(2):12394-12406. doi: 10.1080/21655979.2021.2005217. Bioengineered. 2021. PMID: 34787051 Free PMC article.
-
Deubiquitinating Enzymes: A Critical Regulator of Mitosis.Int J Mol Sci. 2019 Nov 28;20(23):5997. doi: 10.3390/ijms20235997. Int J Mol Sci. 2019. PMID: 31795161 Free PMC article. Review.
References
-
- Ciechanover A. Stanhill A (2014) The complexity of recognition of ubiquitinated substrates by the 26S proteasome. Biochim Biophys Acta. 1843;1:86–96. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

