The tRNA Elbow in Structure, Recognition and Evolution
- PMID: 26771646
- PMCID: PMC4810234
- DOI: 10.3390/life6010003
The tRNA Elbow in Structure, Recognition and Evolution
Abstract
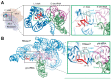
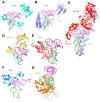
Prominent in the L-shaped three-dimensional structure of tRNAs is the "elbow" where their two orthogonal helical stacks meet. It has a conserved structure arising from the interaction of the terminal loops of the D- and T-stem-loops, and presents to solution a flat face of a tertiary base pair between the D- and T-loops. In addition to the ribosome, which interacts with the elbow in all three of its tRNA binding sites, several cellular RNAs and many proteins are known to recognize the elbow. At least three classes of non-coding RNAs, namely 23S rRNA, ribonuclease P, and the T-box riboswitches, recognize the tRNA elbow employing an identical structural motif consisting of two interdigitated T-loops. In contrast, structural solutions to tRNA-elbow recognition by proteins are varied. Some enzymes responsible for post-transcriptional tRNA modification even disrupt the elbow structure in order to access their substrate nucleotides. The evolutionary origin of the elbow is mysterious, but, because it does not explicitly participate in the flow of genetic information, it has been proposed to be a late innovation. Regardless, it is biologically essential. Even some viruses that hijack the cellular machinery using tRNA decoys have convergently evolved near-perfect mimics of the tRNA elbow.
Keywords: RNA structure; T-loop; base stacking; convergent evolution; ribosome; tRNA elbow.
Figures



Similar articles
-
Co-crystal structure of a T-box riboswitch stem I domain in complex with its cognate tRNA.Nature. 2013 Aug 15;500(7462):363-6. doi: 10.1038/nature12440. Epub 2013 Jul 28. Nature. 2013. PMID: 23892783 Free PMC article.
-
A universal RNA structural motif docking the elbow of tRNA in the ribosome, RNAse P and T-box leaders.Nucleic Acids Res. 2013 May 1;41(10):5494-502. doi: 10.1093/nar/gkt219. Epub 2013 Apr 10. Nucleic Acids Res. 2013. PMID: 23580544 Free PMC article.
-
Post-Transcriptional Modifications of Conserved Nucleotides in the T-Loop of tRNA: A Tale of Functional Convergent Evolution.Genes (Basel). 2021 Jan 22;12(2):140. doi: 10.3390/genes12020140. Genes (Basel). 2021. PMID: 33499018 Free PMC article. Review.
-
Recognition of the T stem-loop of a pre-tRNA substrate by the ribozyme from Bacillus subtilis ribonuclease P.Biochemistry. 1997 May 27;36(21):6317-25. doi: 10.1021/bi970115o. Biochemistry. 1997. PMID: 9174346
-
Trying on tRNA for Size: RNase P and the T-box Riboswitch as Molecular Rulers.Biomolecules. 2016 Apr 1;6(2):18. doi: 10.3390/biom6020018. Biomolecules. 2016. PMID: 27043647 Free PMC article. Review.
Cited by
-
Tuning tRNAs for improved translation.Front Genet. 2024 Jun 25;15:1436860. doi: 10.3389/fgene.2024.1436860. eCollection 2024. Front Genet. 2024. PMID: 38983271 Free PMC article. Review.
-
Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA.Nat Commun. 2022 Aug 30;13(1):5100. doi: 10.1038/s41467-022-32544-1. Nat Commun. 2022. PMID: 36042193 Free PMC article.
-
The Thermus thermophilus DEAD-box protein Hera is a general RNA binding protein and plays a key role in tRNA metabolism.RNA. 2020 Nov;26(11):1557-1574. doi: 10.1261/rna.075580.120. Epub 2020 Jul 15. RNA. 2020. PMID: 32669294 Free PMC article.
-
Molecular basis of human nuclear and mitochondrial tRNA 3' processing.Nat Struct Mol Biol. 2025 Apr;32(4):613-624. doi: 10.1038/s41594-024-01445-w. Epub 2025 Jan 2. Nat Struct Mol Biol. 2025. PMID: 39747487 Free PMC article.
-
Bacillus subtilis, the model Gram-positive bacterium: 20 years of annotation refinement.Microb Biotechnol. 2018 Jan;11(1):3-17. doi: 10.1111/1751-7915.13043. Microb Biotechnol. 2018. PMID: 29280348 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

