Genomic landscapes of endogenous retroviruses unveil intricate genetics of conventional and genetically-engineered laboratory mouse strains
- PMID: 26779669
- PMCID: PMC4823159
- DOI: 10.1016/j.yexmp.2016.01.005
Genomic landscapes of endogenous retroviruses unveil intricate genetics of conventional and genetically-engineered laboratory mouse strains
Abstract
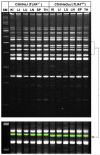
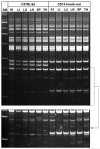
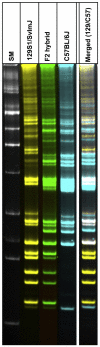
Laboratory strains of mice, both conventional and genetically engineered, have been introduced as critical components of a broad range of studies investigating normal and disease biology. Currently, the genetic identity of laboratory mice is primarily confirmed by surveying polymorphisms in selected sets of "conventional" genes and/or microsatellites in the absence of a single completely sequenced mouse genome. First, we examined variations in the genomic landscapes of transposable repetitive elements, named the TREome, in conventional and genetically engineered mouse strains using murine leukemia virus-type endogenous retroviruses (MLV-ERVs) as a probe. A survey of the genomes from 56 conventional strains revealed strain-specific TREome landscapes, and certain families (e.g., C57BL) of strains were discernible with defined patterns. Interestingly, the TREome landscapes of C3H/HeJ (toll-like receptor-4 [TLR4] mutant) inbred mice were different from its control C3H/HeOuJ (TLR4 wild-type) strain. In addition, a CD14 knock-out strain had a distinct TREome landscape compared to its control/backcross C57BL/6J strain. Second, an examination of superantigen (SAg, a "TREome gene") coding sequences of mouse mammary tumor virus-type ERVs in the genomes of the 46 conventional strains revealed a high diversity, suggesting a potential role of SAgs in strain-specific immune phenotypes. The findings from this study indicate that unexplored and intricate genomic variations exist in laboratory mouse strains, both conventional and genetically engineered. The TREome-based high-resolution genetics surveillance system for laboratory mice would contribute to efficient study design with quality control and accurate data interpretation. This genetics system can be easily adapted to other species ranging from plants to humans.
Keywords: Genetic surveillance; Genome complexity; TREome gene profile; TREome landscape; Transposable repetitive elements (TREome).
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures

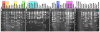


Similar articles
-
Highly Variable Genomic Landscape of Endogenous Retroviruses in the C57BL/6J Inbred Strain, Depending on Individual Mouse, Gender, Organ Type, and Organ Location.Int J Genomics. 2017;2017:3152410. doi: 10.1155/2017/3152410. Epub 2017 Aug 29. Int J Genomics. 2017. PMID: 28951865 Free PMC article.
-
Distribution of endogenous gammaretroviruses and variants of the Fv1 restriction gene in individual mouse strains and strain subgroups.PLoS One. 2019 Jul 10;14(7):e0219576. doi: 10.1371/journal.pone.0219576. eCollection 2019. PLoS One. 2019. PMID: 31291374 Free PMC article.
-
Intracisternal A particle genes: Distribution in the mouse genome, active subtypes, and potential roles as species-specific mediators of susceptibility to cancer.Mol Carcinog. 2010 Jan;49(1):54-67. doi: 10.1002/mc.20576. Mol Carcinog. 2010. PMID: 20025072
-
Origins of the endogenous and infectious laboratory mouse gammaretroviruses.Viruses. 2014 Dec 26;7(1):1-26. doi: 10.3390/v7010001. Viruses. 2014. PMID: 25549291 Free PMC article. Review.
-
Complete overview of protein-inactivating sequence variations in 36 sequenced mouse inbred strains.Proc Natl Acad Sci U S A. 2017 Aug 22;114(34):9158-9163. doi: 10.1073/pnas.1706168114. Epub 2017 Aug 7. Proc Natl Acad Sci U S A. 2017. PMID: 28784771 Free PMC article. Review.
References
-
- Acha-Orbea H, MacDonald HR. Superantigens of mouse mammary tumor virus. Annu. Rev. Immunol. 1995;13:459–486. - PubMed
-
- Bentvelzen P. Immunosuppression by the MMTV superantigen? Immunol. Today. 1992;13:77. - PubMed
-
- Beutler B. Tlr4: central component of the sole mammalian LPS sensor. Curr. Opin. Immunol. 2000;12:20–26. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

