The intolerance to functional genetic variation of protein domains predicts the localization of pathogenic mutations within genes
- PMID: 26781712
- PMCID: PMC4717634
- DOI: 10.1186/s13059-016-0869-4
The intolerance to functional genetic variation of protein domains predicts the localization of pathogenic mutations within genes
Abstract
Ranking human genes based on their tolerance to functional genetic variation can greatly facilitate patient genome interpretation. It is well established, however, that different parts of proteins can have different functions, suggesting that it will ultimately be more informative to focus attention on functionally distinct portions of genes. Here we evaluate the intolerance of genic sub-regions using two biological sub-region classifications. We show that the intolerance scores of these sub-regions significantly correlate with reported pathogenic mutations. This observation extends the utility of intolerance scores to indicating where pathogenic mutations are mostly likely to fall within genes.
Figures

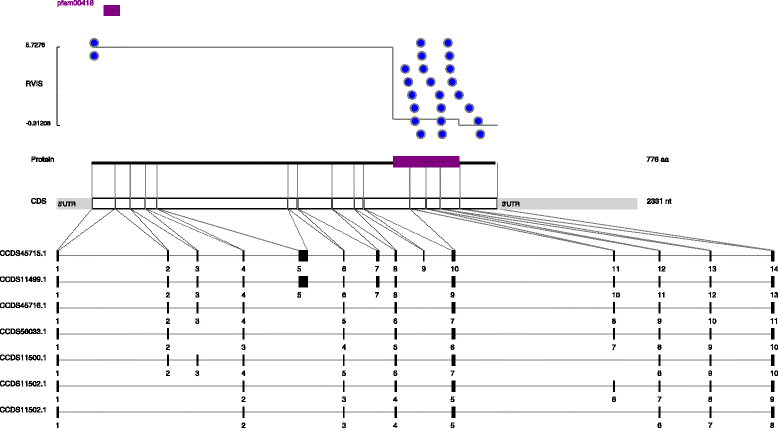
References
Publication types
MeSH terms
Grants and funding
- F31 NS092362/NS/NINDS NIH HHS/United States
- RC2 HL102923/HL/NHLBI NIH HHS/United States
- UC2 HL102926/HL/NHLBI NIH HHS/United States
- UC2 HL103010/HL/NHLBI NIH HHS/United States
- U01NS077303/NS/NINDS NIH HHS/United States
- RC2 HL102926/HL/NHLBI NIH HHS/United States
- RC2 HL102924/HL/NHLBI NIH HHS/United States
- U01 NS077303/NS/NINDS NIH HHS/United States
- T32 GM071340/GM/NIGMS NIH HHS/United States
- UC2 HL102923/HL/NHLBI NIH HHS/United States
- UC2 HL102924/HL/NHLBI NIH HHS/United States
- RC2 HL103010/HL/NHLBI NIH HHS/United States
- RC2 HL102925/HL/NHLBI NIH HHS/United States
- UC2 HL102925/HL/NHLBI NIH HHS/United States
- F31NS092362/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

