Tissue-Specific and Genetic Regulation of Insulin Sensitivity-Associated Transcripts in African Americans
- PMID: 26789776
- PMCID: PMC4880154
- DOI: 10.1210/jc.2015-3336
Tissue-Specific and Genetic Regulation of Insulin Sensitivity-Associated Transcripts in African Americans
Abstract
Integrative multiomics analyses of adipose and muscle tissue transcripts, S, and genotypes revealed novel genetic regulatory mechanisms of insulin resistance in African Americans.
Figures
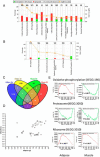

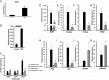
References
-
- Johnson AM, Olefsky JM. The origins and drivers of insulin resistance. Cell. 2013;152(4):673–684. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

