Correlation of the lung microbiota with metabolic profiles in bronchoalveolar lavage fluid in HIV infection
- PMID: 26792212
- PMCID: PMC4721204
- DOI: 10.1186/s40168-016-0147-4
Correlation of the lung microbiota with metabolic profiles in bronchoalveolar lavage fluid in HIV infection
Abstract
Background: While 16S ribosomal RNA (rRNA) sequencing has been used to characterize the lung's bacterial microbiota in human immunodeficiency virus (HIV)-infected individuals, taxonomic studies provide limited information on bacterial function and impact on the host. Metabolic profiles can provide functional information on host-microbe interactions in the lungs. We investigated the relationship between the respiratory microbiota and metabolic profiles in the bronchoalveolar lavage fluid of HIV-infected and HIV-uninfected outpatients.
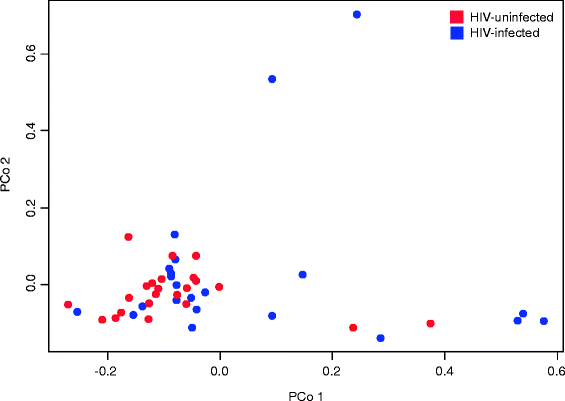
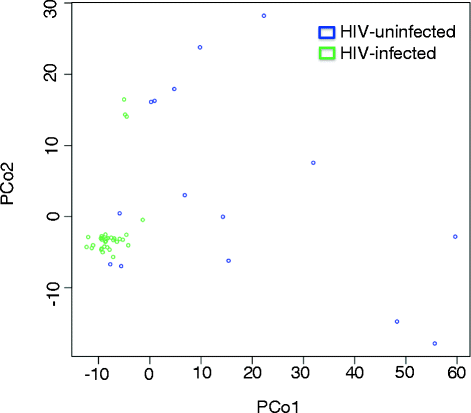
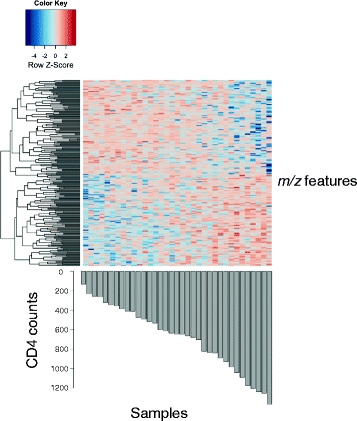
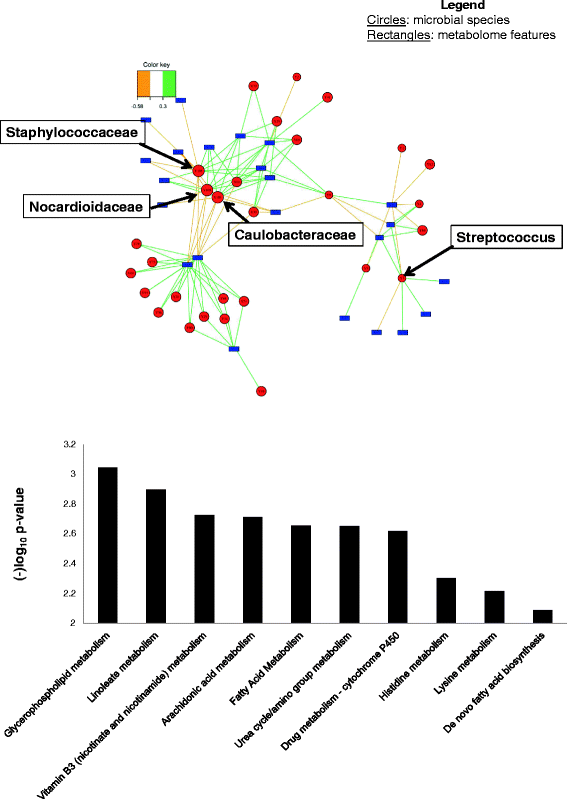
Results: Targeted sequencing of the 16S rRNA gene was used to analyze the bacterial community structure and liquid chromatography-high-resolution mass spectrometry was used to detect features in bronchoalveolar lavage fluid. Global integration of all metabolic features with microbial species was done using sparse partial least squares regression. Thirty-nine HIV-infected subjects and 20 HIV-uninfected controls without acute respiratory symptoms were enrolled. Twelve mass-to-charge ratio (m/z) features from C18 analysis were significantly different between HIV-infected individuals and controls (false discovery rate (FDR) = 0.2); another 79 features were identified by network analysis. Further metabolite analysis demonstrated that four features were significantly overrepresented in the bronchoalveolar lavage (BAL) fluid of HIV-infected individuals compared to HIV-uninfected, including cystine, two complex carbohydrates, and 3,5-dibromo-L-tyrosine. There were 231 m/z features significantly associated with peripheral blood CD4 cell counts identified using sparse partial least squares regression (sPLS) at a variable importance on projection (VIP) threshold of 2. Twenty-five percent of these 91 m/z features were associated with various microbial species. Bacteria from families Caulobacteraceae, Staphylococcaceae, Nocardioidaceae, and genus Streptococcus were associated with the greatest number of features. Glycerophospholipid and lineolate pathways correlated with these bacteria.
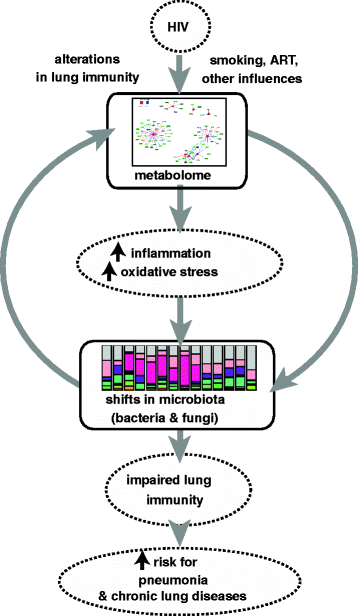
Conclusions: In bronchoalveolar lavage fluid, specific metabolic profiles correlated with bacterial organisms known to play a role in the pathogenesis of pneumonia in HIV-infected individuals. These findings suggest that microbial communities and their interactions with the host may have functional metabolic impact in the lung.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
- K24 HL123342/HL/NHLBI NIH HHS/United States
- U01 AI035042/AI/NIAID NIH HHS/United States
- UO1-HD-32632/HD/NICHD NIH HHS/United States
- U01 AI037984/AI/NIAID NIH HHS/United States
- UL1TR000124/TR/NCATS NIH HHS/United States
- U01 HL098962/HL/NHLBI NIH HHS/United States
- UO1-AI-34994/AI/NIAID NIH HHS/United States
- UO1-AI-34989/AI/NIAID NIH HHS/United States
- U01 AI031834/AI/NIAID NIH HHS/United States
- UO1-AI-35042/AI/NIAID NIH HHS/United States
- UO1-AI-37984/AI/NIAID NIH HHS/United States
- UL1-TR000004/TR/NCATS NIH HHS/United States
- U01 AI035004/AI/NIAID NIH HHS/United States
- U01-AI35040/AI/NIAID NIH HHS/United States
- NIH HL113451/HL/NHLBI NIH HHS/United States
- K24HL123342/HL/NHLBI NIH HHS/United States
- UO1-AI-35040/AI/NIAID NIH HHS/United States
- UL1 RR024131/RR/NCRR NIH HHS/United States
- HL087713/HL/NHLBI NIH HHS/United States
- U01 AI034989/AI/NIAID NIH HHS/United States
- UL1 TR000124/TR/NCATS NIH HHS/United States
- U01 AI037613/AI/NIAID NIH HHS/United States
- UO1-AI-35041/AI/NIAID NIH HHS/United States
- M01 RR000722/RR/NCRR NIH HHS/United States
- UO1-AI-35004/AI/NIAID NIH HHS/United States
- S10 OD018006/OD/NIH HHS/United States
- U01 AI035041/AI/NIAID NIH HHS/United States
- UM1 AI035043/AI/NIAID NIH HHS/United States
- UO1-AI-34993/AI/NIAID NIH HHS/United States
- U01 AI034994/AI/NIAID NIH HHS/United States
- K24 HL087713/HL/NHLBI NIH HHS/United States
- UL1 TR000424/TR/NCATS NIH HHS/United States
- R01 HL090339/HL/NHLBI NIH HHS/United States
- U01-AI35041/AI/NIAID NIH HHS/United States
- U01 AI035043/AI/NIAID NIH HHS/United States
- NIH R01 HL090339/HL/NHLBI NIH HHS/United States
- UO1-AI-35039/AI/NIAID NIH HHS/United States
- UO1-AI-42590/AI/NIAID NIH HHS/United States
- U01 AI035040/AI/NIAID NIH HHS/United States
- U01 AI034993/AI/NIAID NIH HHS/United States
- UL1-TR000424/TR/NCATS NIH HHS/United States
- P20 HL113451/HL/NHLBI NIH HHS/United States
- 5-MO1-RR-00722/RR/NCRR NIH HHS/United States
- UO1-AI-31834/AI/NIAID NIH HHS/United States
- UL1 TR000004/TR/NCATS NIH HHS/United States
- U01 AI103408/AI/NIAID NIH HHS/United States
- U01 AI035039/AI/NIAID NIH HHS/United States
- UO1-AI-35043/AI/NIAID NIH HHS/United States
- UO1-AI-37613/AI/NIAID NIH HHS/United States
- UM1-AI35043/AI/NIAID NIH HHS/United States
- U01 HD032632/HD/NICHD NIH HHS/United States
- U01 AI042590/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

