Clinical evaluation of panel testing by next-generation sequencing (NGS) for gene mutations in myeloid neoplasms
- PMID: 26796102
- PMCID: PMC4722624
- DOI: 10.1186/s13000-016-0456-8
Clinical evaluation of panel testing by next-generation sequencing (NGS) for gene mutations in myeloid neoplasms
Abstract
Background: Genomic techniques in recent years have allowed the identification of many mutated genes important in the pathogenesis of acute myeloid leukemia (AML). Together with cytogenetic aberrations, these gene mutations are powerful prognostic markers in AML and can be used to guide patient management, for example selection of optimal post-remission therapy. The mutated genes also hold promise as therapeutic targets themselves. We evaluated the applicability of a gene panel for the detection of AML mutations in a diagnostic molecular pathology laboratory.
Methods: Fifty patient samples comprising 46 AML and 4 other myeloid neoplasms were accrued for the study. They consisted of 19 males and 31 females at a median age of 60 years (range: 18-88 years). A total of 54 genes (full coding exons of 15 genes and exonic hotspots of 39 genes) were targeted by 568 amplicons that ranged from 225 to 275 bp. The combined coverage was 141 kb in sequence length. Amplicon libraries were prepared by TruSight myeloid sequencing panel (Illumina, CA) and paired-end sequencing runs were performed on a MiSeq (Illumina) genome sequencer. Sequences obtained were analyzed by in-house bioinformatics pipeline, namely BWA-MEM, Samtools, GATK, Pindel, Ensembl Variant Effect Predictor and a novel algorithm ITDseek.
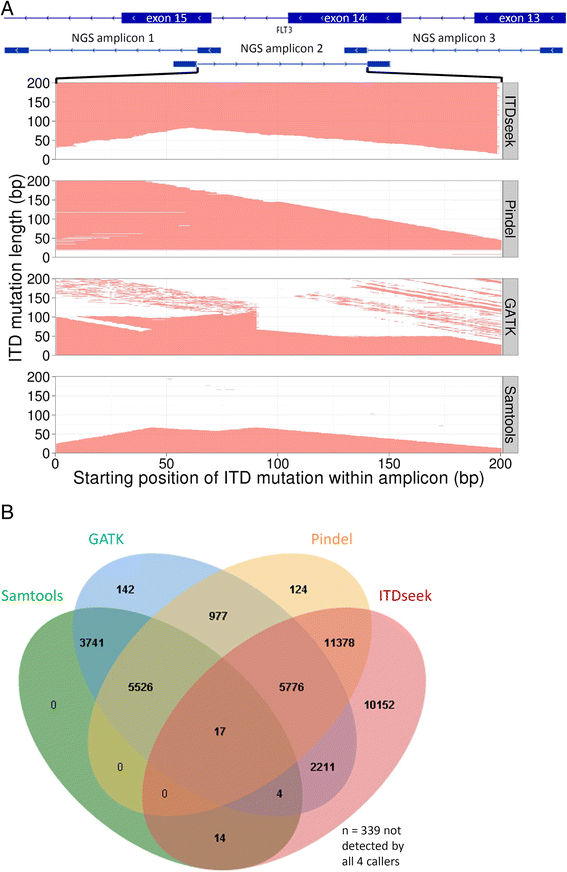
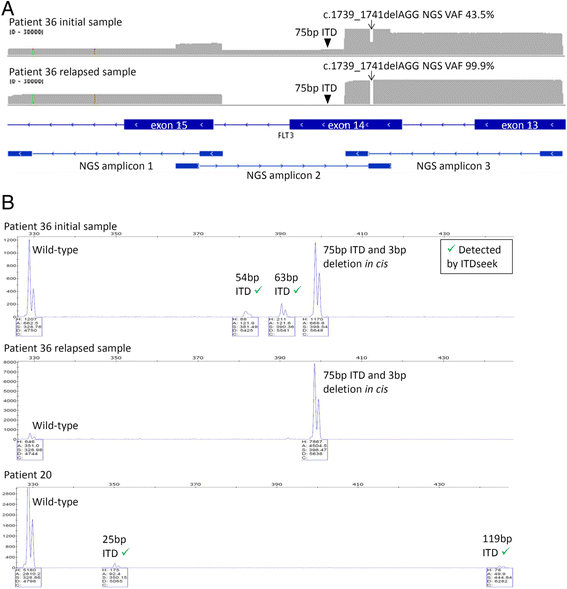
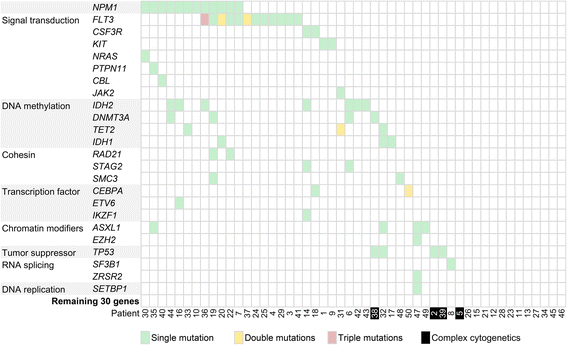
Results: The mean count of sequencing reads obtained per sample was 3.81 million and the mean sequencing depth was over 3000X. Seventy-seven mutations in 24 genes were detected in 37 of 50 samples (74 %). On average, 2 mutations (range 1-5) were detected per positive sample. TP53 gene mutations were found in 3 out of 4 patients with complex and unfavorable cytogenetics. Comparing NGS results with that of conventional molecular testing showed a concordance rate of 95.5 %. After further resolution and application of a novel bioinformatics algorithm ITDseek to aid the detection of FLT3 internal tandem duplication (ITD), the concordance rate was revised to 98.2 %.
Conclusions: Gene panel testing by NGS approach was applicable for sensitive and accurate detection of actionable AML gene mutations in the clinical laboratory to individualize patient management. A novel algorithm ITDseek was presented that improved the detection of FLT3-ITD of varying length, position and at low allelic burden.
Figures
References
-
- Grimwade D, Hills RK, Moorman AV, Walker H, Chatters S, Goldstone AH, et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood. 2010;116(3):354–65. doi: 10.1182/blood-2009-11-254441. - DOI - PubMed
-
- Dohner H, Estey EH, Amadori S, Appelbaum FR, Buchner T, Burnett AK, et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood. 2010;115(3):453–74. doi: 10.1182/blood-2009-07-235358. - DOI - PubMed
-
- Taskesen E, Bullinger L, Corbacioglu A, Sanders MA, Erpelinck CA, Wouters BJ, et al. Prognostic impact, concurrent genetic mutations, and gene expression features of AML with CEBPA mutations in a cohort of 1182 cytogenetically normal AML patients: further evidence for CEBPA double mutant AML as a distinctive disease entity. Blood. 2011;117(8):2469–75. doi: 10.1182/blood-2010-09-307280. - DOI - PubMed
-
- Wouters BJ, Lowenberg B, Erpelinck-Verschueren CA, van Putten WL, Valk PJ, Delwel R. Double CEBPA mutations, but not single CEBPA mutations, define a subgroup of acute myeloid leukemia with a distinctive gene expression profile that is uniquely associated with a favorable outcome. Blood. 2009;113(13):3088–91. doi: 10.1182/blood-2008-09-179895. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

