Differential Expression of Genes Involved in Host Recognition, Attachment, and Degradation in the Mycoparasite Tolypocladium ophioglossoides
- PMID: 26801645
- PMCID: PMC4777134
- DOI: 10.1534/g3.116.027045
Differential Expression of Genes Involved in Host Recognition, Attachment, and Degradation in the Mycoparasite Tolypocladium ophioglossoides
Abstract
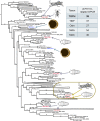
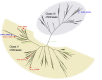
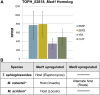
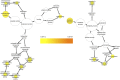
The ability of a fungus to infect novel hosts is dependent on changes in gene content, expression, or regulation. Examining gene expression under simulated host conditions can explore which genes may contribute to host jumping. Insect pathogenesis is the inferred ancestral character state for species of Tolypocladium, however several species are parasites of truffles, including Tolypocladium ophioglossoides. To identify potentially crucial genes in this interkingdom host switch, T. ophioglossoides was grown on four media conditions: media containing the inner and outer portions of its natural host (truffles of Elaphomyces), cuticles from an ancestral host (beetle), and a rich medium (Yeast Malt). Through high-throughput RNASeq of mRNA from these conditions, many differentially expressed genes were identified in the experiment. These included PTH11-related G-protein-coupled receptors (GPCRs) hypothesized to be involved in host recognition, and also found to be upregulated in insect pathogens. A divergent chitinase with a signal peptide was also found to be highly upregulated on media containing truffle tissue, suggesting an exogenous degradative activity in the presence of the truffle host. The adhesin gene, Mad1, was highly expressed on truffle media as well. A BiNGO analysis of overrepresented GO terms from genes expressed during each growth condition found that genes involved in redox reactions and transmembrane transport were the most overrepresented during T. ophioglossoides growth on truffle media, suggesting their importance in growth on fungal tissue as compared to other hosts and environments. Genes involved in secondary metabolism were most highly expressed during growth on insect tissue, suggesting that their products may not be necessary during parasitism of Elaphomyces. This study provides clues into understanding genetic mechanisms underlying the transition from insect to truffle parasitism.
Keywords: Elaphomyces; PTH11 GPCR; RNA-Seq; adhesin; chitinase.
Copyright © 2016 Quandt et al.
Figures
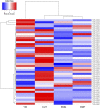
Similar articles
-
Harnessing the power of phylogenomics to disentangle the directionality and signatures of interkingdom host jumping in the parasitic fungal genus Tolypocladium.Mycologia. 2018 Jan-Feb;110(1):104-117. doi: 10.1080/00275514.2018.1442618. Mycologia. 2018. PMID: 29863984
-
The genome of the truffle-parasite Tolypocladium ophioglossoides and the evolution of antifungal peptaibiotics.BMC Genomics. 2015 Jul 28;16(1):553. doi: 10.1186/s12864-015-1777-9. BMC Genomics. 2015. PMID: 26215153 Free PMC article.
-
Natural infection of Chiromyzinae larvae (Diptera: Stratiomyidae) in southern Chile by Tolypocladium valdiviae sp. nov.Fungal Biol. 2023 Jan-Feb;127(1-2):845-853. doi: 10.1016/j.funbio.2022.12.004. Epub 2022 Dec 24. Fungal Biol. 2023. PMID: 36746556
-
Genome-wide comparative analysis of putative Pth11-related G protein-coupled receptors in fungi belonging to Pezizomycotina.BMC Microbiol. 2017 Jul 25;17(1):166. doi: 10.1186/s12866-017-1076-5. BMC Microbiol. 2017. PMID: 28743231 Free PMC article.
-
Truffles: much more than a prized and local fungal delicacy.FEMS Microbiol Lett. 2006 Jul;260(1):1-8. doi: 10.1111/j.1574-6968.2006.00252.x. FEMS Microbiol Lett. 2006. PMID: 16790011 Review.
Cited by
-
Comprehensive Review of Tolypocladium and Description of a Novel Lineage from Southwest China.Pathogens. 2021 Oct 27;10(11):1389. doi: 10.3390/pathogens10111389. Pathogens. 2021. PMID: 34832545 Free PMC article.
-
Tolypocladamide H and the Proposed Tolypocladamide NRPS in Tolypocladium Species.J Nat Prod. 2022 May 27;85(5):1363-1373. doi: 10.1021/acs.jnatprod.2c00153. Epub 2022 May 2. J Nat Prod. 2022. PMID: 35500108 Free PMC article. Review.
-
The mycoparasitic fungus Clonostachys rosea responds with both common and specific gene expression during interspecific interactions with fungal prey.Evol Appl. 2018 Mar 14;11(6):931-949. doi: 10.1111/eva.12609. eCollection 2018 Jul. Evol Appl. 2018. PMID: 29928301 Free PMC article.
-
Comparative genomics reveals dynamic genome evolution in host specialist ectomycorrhizal fungi.New Phytol. 2021 Apr;230(2):774-792. doi: 10.1111/nph.17160. Epub 2021 Feb 6. New Phytol. 2021. PMID: 33355923 Free PMC article.
-
Transcriptomics Reveals the Putative Mycoparasitic Strategy of the Mushroom Entoloma abortivum on Species of the Mushroom Genus Armillaria.mSystems. 2021 Oct 26;6(5):e0054421. doi: 10.1128/mSystems.00544-21. Epub 2021 Oct 12. mSystems. 2021. PMID: 34636668 Free PMC article.
References
-
- Andersen S. O., 1980. Cuticular Sclerotization, pp. 185–215 in Cuticle Techniques in Arthropods, edited by Miller T. A. Springer-Verlag, New York.
-
- Chugh J. K., Wallace B. A., 2001. Peptaibols: models for ion channels. Biochem. Soc. Trans. 29: 565–570. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
