Interactions of C4 Subtype Metabolic Activities and Transport in Maize Are Revealed through the Characterization of DCT2 Mutants
- PMID: 26813621
- PMCID: PMC4790864
- DOI: 10.1105/tpc.15.00497
Interactions of C4 Subtype Metabolic Activities and Transport in Maize Are Revealed through the Characterization of DCT2 Mutants
Abstract
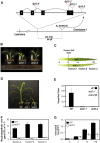
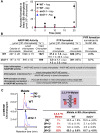
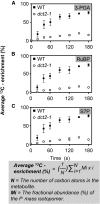
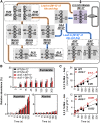
C4 photosynthesis in grasses requires the coordinated movement of metabolites through two specialized leaf cell types, mesophyll (M) and bundle sheath (BS), to concentrate CO2 around Rubisco. Despite the importance of transporters in this process, few have been identified or rigorously characterized. In maize (Zea mays), DCT2 has been proposed to function as a plastid-localized malate transporter and is preferentially expressed in BS cells. Here, we characterized the role of DCT2 in maize leaves using Activator-tagged mutant alleles. Our results indicate that DCT2 enables the transport of malate into the BS chloroplast. Isotopic labeling experiments show that the loss of DCT2 results in markedly different metabolic network operation and dramatically reduced biomass production. In the absence of a functioning malate shuttle, dct2 lines survive through the enhanced use of the phosphoenolpyruvate carboxykinase carbon shuttle pathway that in wild-type maize accounts for ∼ 25% of the photosynthetic activity. The results emphasize the importance of malate transport during C4 photosynthesis, define the role of a primary malate transporter in BS cells, and support a model for carbon exchange between BS and M cells in maize.
© 2016 American Society of Plant Biologists. All rights reserved.
Figures




Similar articles
-
Photosynthetic flexibility in maize exposed to salinity and shade.J Exp Bot. 2014 Jul;65(13):3715-24. doi: 10.1093/jxb/eru130. Epub 2014 Apr 1. J Exp Bot. 2014. PMID: 24692650 Free PMC article.
-
The operation of two decarboxylases, transamination, and partitioning of C4 metabolic processes between mesophyll and bundle sheath cells allows light capture to be balanced for the maize C4 pathway.Plant Physiol. 2014 Jan;164(1):466-80. doi: 10.1104/pp.113.228221. Epub 2013 Nov 19. Plant Physiol. 2014. PMID: 24254314 Free PMC article.
-
Functional analysis of corn husk photosynthesis.Plant Physiol. 2011 Jun;156(2):503-13. doi: 10.1104/pp.111.176495. Epub 2011 Apr 21. Plant Physiol. 2011. PMID: 21511990 Free PMC article.
-
Strategies for engineering a two-celled C(4) photosynthetic pathway into rice.J Exp Bot. 2011 May;62(9):3001-10. doi: 10.1093/jxb/err022. Epub 2011 Feb 18. J Exp Bot. 2011. PMID: 21335436 Review.
-
Strategies for improving C4 photosynthesis.Curr Opin Plant Biol. 2016 Jun;31:125-34. doi: 10.1016/j.pbi.2016.04.003. Epub 2016 Apr 27. Curr Opin Plant Biol. 2016. PMID: 27127850 Review.
Cited by
-
Advances in metabolic flux analysis toward genome-scale profiling of higher organisms.Biosci Rep. 2018 Nov 23;38(6):BSR20170224. doi: 10.1042/BSR20170224. Print 2018 Dec 21. Biosci Rep. 2018. PMID: 30341247 Free PMC article. Review.
-
Metabolite pools and carbon flow during C4 photosynthesis in maize: 13CO2 labeling kinetics and cell type fractionation.J Exp Bot. 2017 Jan;68(2):283-298. doi: 10.1093/jxb/erw414. Epub 2016 Nov 7. J Exp Bot. 2017. PMID: 27834209 Free PMC article.
-
A dominant role of transcriptional regulation during the evolution of C4 photosynthesis in Flaveria species.Nat Commun. 2025 Feb 14;16(1):1643. doi: 10.1038/s41467-025-56901-y. Nat Commun. 2025. PMID: 39952962 Free PMC article.
-
13CO2 labeling kinetics in maize reveal impaired efficiency of C4 photosynthesis under low irradiance.Plant Physiol. 2022 Aug 29;190(1):280-304. doi: 10.1093/plphys/kiac306. Plant Physiol. 2022. PMID: 35751609 Free PMC article.
-
Accelerated remodeling of the mesophyll-bundle sheath interface in the maize C4 cycle mutant leaves.Sci Rep. 2022 Mar 23;12(1):5057. doi: 10.1038/s41598-022-09135-7. Sci Rep. 2022. PMID: 35322159 Free PMC article.
References
-
- Becker T.W., Carrayol E., Hirel B. (2000). Glutamine synthetase and glutamate dehydrogenase isoforms in maize leaves: localization, relative proportion and their role in ammonium assimilation or nitrogen transport. Planta 211: 800–806. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

