An essential receptor for adeno-associated virus infection
- PMID: 26814968
- PMCID: PMC4962915
- DOI: 10.1038/nature16465
An essential receptor for adeno-associated virus infection
Erratum in
-
Corrigendum: An essential receptor for adeno-associated virus infection.Nature. 2016 Nov 17;539(7629):456. doi: 10.1038/nature19835. Epub 2016 Sep 28. Nature. 2016. PMID: 27680708 No abstract available.
Abstract
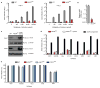
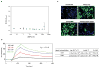
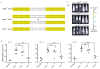
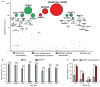
Adeno-associated virus (AAV) vectors are currently the leading candidates for virus-based gene therapies because of their broad tissue tropism, non-pathogenic nature and low immunogenicity. They have been successfully used in clinical trials to treat hereditary diseases such as haemophilia B (ref. 2), and have been approved for treatment of lipoprotein lipase deficiency in Europe. Considerable efforts have been made to engineer AAV variants with novel and biomedically valuable cell tropisms to allow efficacious systemic administration, yet basic aspects of AAV cellular entry are still poorly understood. In particular, the protein receptor(s) required for AAV entry after cell attachment remains unknown. Here we use an unbiased genetic screen to identify proteins essential for AAV serotype 2 (AAV2) infection in a haploid human cell line. The most significantly enriched gene of the screen encodes a previously uncharacterized type I transmembrane protein, KIAA0319L (denoted hereafter as AAV receptor (AAVR)). We characterize AAVR as a protein capable of rapid endocytosis from the plasma membrane and trafficking to the trans-Golgi network. We show that AAVR directly binds to AAV2 particles, and that anti-AAVR antibodies efficiently block AAV2 infection. Moreover, genetic ablation of AAVR renders a wide range of mammalian cell types highly resistant to AAV2 infection. Notably, AAVR serves as a critical host factor for all tested AAV serotypes. The importance of AAVR for in vivo gene delivery is further highlighted by the robust resistance of Aavr(-/-) (also known as Au040320(-/-) and Kiaa0319l(-/-)) mice to AAV infection. Collectively, our data indicate that AAVR is a universal receptor involved in AAV infection.
Conflict of interest statement
The authors declare competing financial interests: Stanford university is in the process of filing a patent claim regarding AAVR applications in AAV vector technology (Inventors: S.P., M.S.C., J.E.C., A.S.P., N.L.M. and O.D.).
Figures







Comment in
-
AAVR: A Multi-Serotype Receptor for AAV.Mol Ther. 2016 Apr;24(4):663-6. doi: 10.1038/mt.2016.49. Mol Ther. 2016. PMID: 27081719 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

