Environmental Association Analyses Identify Candidates for Abiotic Stress Tolerance in Glycine soja, the Wild Progenitor of Cultivated Soybeans
- PMID: 26818076
- PMCID: PMC4825654
- DOI: 10.1534/g3.116.026914
Environmental Association Analyses Identify Candidates for Abiotic Stress Tolerance in Glycine soja, the Wild Progenitor of Cultivated Soybeans
Abstract
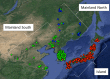
Natural populations across a species range demonstrate population structure owing to neutral processes such as localized origins of mutations and migration limitations. Selection also acts on a subset of loci, contributing to local adaptation. An understanding of the genetic basis of adaptation to local environmental conditions is a fundamental goal in basic biological research. When applied to crop wild relatives, this same research provides the opportunity to identify adaptive genetic variation that may be used to breed for crops better adapted to novel or changing environments. The present study explores an ex situ conservation collection, the USDA germplasm collection, genotyped at 32,416 SNPs to identify population structure and test for associations with bioclimatic and biophysical variables in Glycine soja, the wild progenitor of Glycine max (soybean). Candidate loci were detected that putatively contribute to adaptation to abiotic stresses. The identification of potentially adaptive variants in this ex situ collection may permit a more targeted use of germplasm collections.
Keywords: Glycine soja; crop wild relative; germplasm collections; landscape genomics; population structure; soybean.
Copyright © 2016 Anderson et al.
Figures

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
