UbiSite: incorporating two-layered machine learning method with substrate motifs to predict ubiquitin-conjugation site on lysines
- PMID: 26818456
- PMCID: PMC4895383
- DOI: 10.1186/s12918-015-0246-z
UbiSite: incorporating two-layered machine learning method with substrate motifs to predict ubiquitin-conjugation site on lysines
Abstract
Background: The conjugation of ubiquitin to a substrate protein (protein ubiquitylation), which involves a sequential process--E1 activation, E2 conjugation and E3 ligation, is crucial to the regulation of protein function and activity in eukaryotes. This ubiquitin-conjugation process typically binds the last amino acid of ubiquitin (glycine 76) to a lysine residue of a target protein. The high-throughput of mass spectrometry-based proteomics has stimulated a large-scale identification of ubiquitin-conjugated peptides. Hence, a new web resource, UbiSite, was developed to identify ubiquitin-conjugation site on lysines based on large-scale proteome dataset.
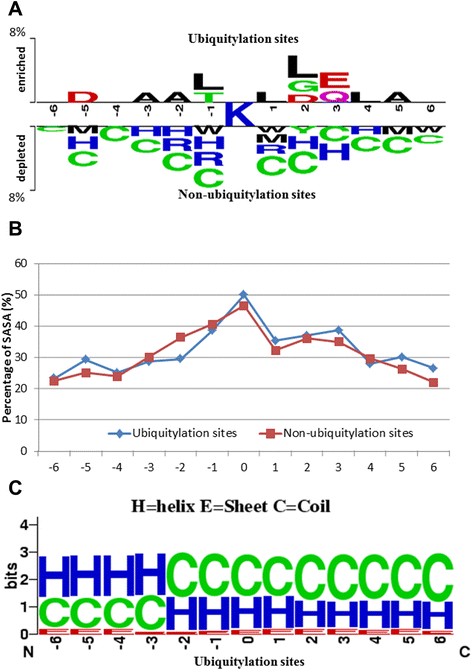
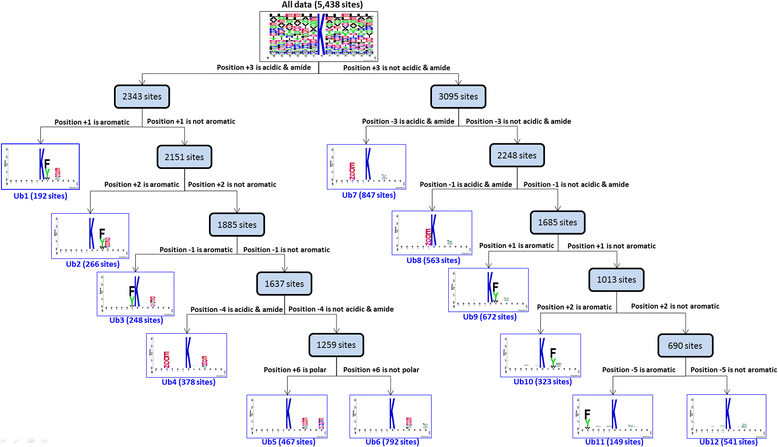
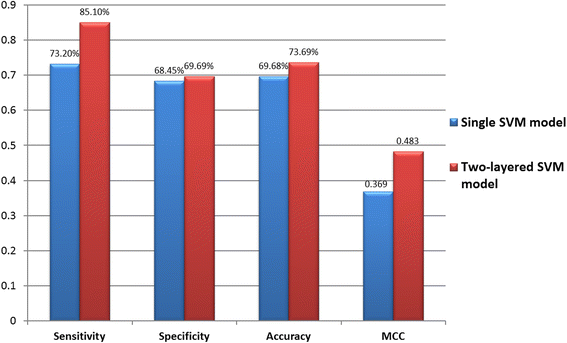
Results: Given a total of 37,647 ubiquitin-conjugated proteins, including 128,026 ubiquitylated peptides, obtained from various resources, this study carries out a large-scale investigation on ubiquitin-conjugation sites based on sequenced and structural characteristics. A TwoSampleLogo reveals that a significant depletion of histidine (H), arginine (R) and cysteine (C) residues around ubiquitylation sites may impact the conjugation of ubiquitins in closed three-dimensional environments. Based on the large-scale ubiquitylation dataset, a motif discovery tool, MDDLogo, has been adopted to characterize the potential substrate motifs for ubiquitin conjugation. Not only are single features such as amino acid composition (AAC), positional weighted matrix (PWM), position-specific scoring matrix (PSSM) and solvent-accessible surface area (SASA) considered, but also the effectiveness of incorporating MDDLogo-identified substrate motifs into a two-layered prediction model is taken into account. Evaluation by five-fold cross-validation showed that PSSM is the best feature in discriminating between ubiquitylation and non-ubiquitylation sites, based on support vector machine (SVM). Additionally, the two-layered SVM model integrating MDDLogo-identified substrate motifs could obtain a promising accuracy and the Matthews Correlation Coefficient (MCC) at 81.06% and 0.586, respectively. Furthermore, the independent testing showed that the two-layered SVM model could outperform other prediction tools, reaching at 85.10% sensitivity, 69.69% specificity, 73.69% accuracy and the 0.483 of MCC value.
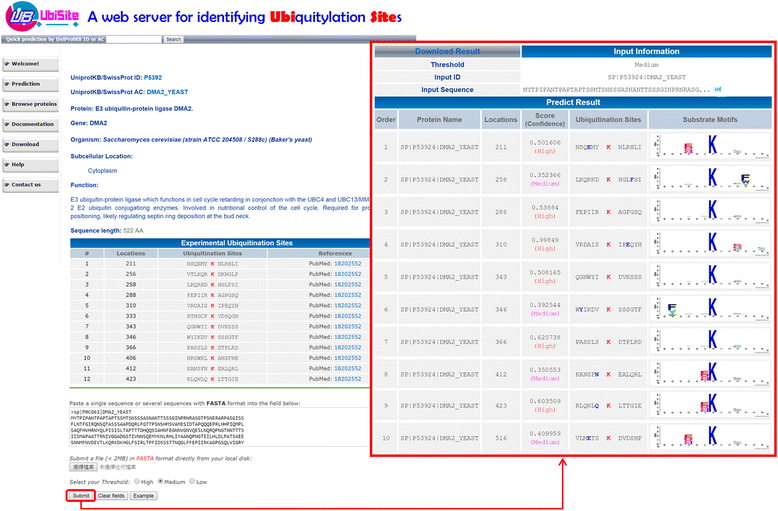
Conclusion: The independent testing result indicated the effectiveness of incorporating MDDLogo-identified motifs into the prediction of ubiquitylation sites. In order to provide meaningful assistance to researchers interested in large-scale ubiquitinome data, the two-layered SVM model has been implemented onto a web-based system (UbiSite), which is freely available at http://csb.cse.yzu.edu.tw/UbiSite/ . Two cases given in the UbiSite provide a demonstration of effective identification of ubiquitylation sites with reference to substrate motifs.
Figures

Similar articles
-
MDD-Palm: Identification of protein S-palmitoylation sites with substrate motifs based on maximal dependence decomposition.PLoS One. 2017 Jun 29;12(6):e0179529. doi: 10.1371/journal.pone.0179529. eCollection 2017. PLoS One. 2017. PMID: 28662047 Free PMC article.
-
A New Scheme to Characterize and Identify Protein Ubiquitination Sites.IEEE/ACM Trans Comput Biol Bioinform. 2017 Mar-Apr;14(2):393-403. doi: 10.1109/TCBB.2016.2520939. Epub 2016 Feb 8. IEEE/ACM Trans Comput Biol Bioinform. 2017. PMID: 26887002
-
Characterization and identification of lysine glutarylation based on intrinsic interdependence between positions in the substrate sites.BMC Bioinformatics. 2019 Feb 4;19(Suppl 13):384. doi: 10.1186/s12859-018-2394-9. BMC Bioinformatics. 2019. PMID: 30717647 Free PMC article.
-
Unraveling the ubiquitin-regulated signaling networks by mass spectrometry-based proteomics.Proteomics. 2013 Feb;13(3-4):526-37. doi: 10.1002/pmic.201200244. Epub 2012 Nov 26. Proteomics. 2013. PMID: 23019148 Review.
-
Advances in characterizing ubiquitylation sites by mass spectrometry.Curr Opin Chem Biol. 2013 Feb;17(1):49-58. doi: 10.1016/j.cbpa.2012.12.009. Epub 2013 Jan 5. Curr Opin Chem Biol. 2013. PMID: 23298953 Review.
Cited by
-
UbiComb: A Hybrid Deep Learning Model for Predicting Plant-Specific Protein Ubiquitylation Sites.Genes (Basel). 2021 May 11;12(5):717. doi: 10.3390/genes12050717. Genes (Basel). 2021. PMID: 34064731 Free PMC article.
-
Multi-dimensional feature recognition model based on capsule network for ubiquitination site prediction.PeerJ. 2022 Dec 6;10:e14427. doi: 10.7717/peerj.14427. eCollection 2022. PeerJ. 2022. PMID: 36523471 Free PMC article.
-
Macrophage IRX3 promotes diet-induced obesity and metabolic inflammation.Nat Immunol. 2021 Oct;22(10):1268-1279. doi: 10.1038/s41590-021-01023-y. Epub 2021 Sep 23. Nat Immunol. 2021. PMID: 34556885
-
Mini-review: Recent advances in post-translational modification site prediction based on deep learning.Comput Struct Biotechnol J. 2022 Jun 30;20:3522-3532. doi: 10.1016/j.csbj.2022.06.045. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35860402 Free PMC article. Review.
-
Identification of apolipoprotein using feature selection technique.Sci Rep. 2016 Jul 22;6:30441. doi: 10.1038/srep30441. Sci Rep. 2016. PMID: 27443605 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

