Unbiased Detection of Respiratory Viruses by Use of RNA Sequencing-Based Metagenomics: a Systematic Comparison to a Commercial PCR Panel
- PMID: 26818672
- PMCID: PMC4809917
- DOI: 10.1128/JCM.03060-15
Unbiased Detection of Respiratory Viruses by Use of RNA Sequencing-Based Metagenomics: a Systematic Comparison to a Commercial PCR Panel
Abstract
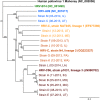
Current infectious disease molecular tests are largely pathogen specific, requiring test selection based on the patient's symptoms. For many syndromes caused by a large number of viral, bacterial, or fungal pathogens, such as respiratory tract infections, this necessitates large panels of tests and has limited yield. In contrast, next-generation sequencing-based metagenomics can be used for unbiased detection of any expected or unexpected pathogen. However, barriers for its diagnostic implementation include incomplete understanding of analytical performance and complexity of sequence data analysis. We compared detection of known respiratory virus-positive (n= 42) and unselected (n= 67) pediatric nasopharyngeal swabs using an RNA sequencing (RNA-seq)-based metagenomics approach and Taxonomer, an ultrarapid, interactive, web-based metagenomics data analysis tool, with an FDA-cleared respiratory virus panel (RVP; GenMark eSensor). Untargeted metagenomics detected 86% of known respiratory virus infections, and additional PCR testing confirmed RVP results for only 2 (33%) of the discordant samples. In unselected samples, untargeted metagenomics had excellent agreement with the RVP (93%). In addition, untargeted metagenomics detected an additional 12 viruses that were either not targeted by the RVP or missed due to highly divergent genome sequences. Normalized viral read counts for untargeted metagenomics correlated with viral burden determined by quantitative PCR and showed high intrarun and interrun reproducibility. Partial or full-length viral genome sequences were generated in 86% of RNA-seq-positive samples, allowing assessment of antiviral resistance, strain-level typing, and phylogenetic relatedness. Overall, untargeted metagenomics had high agreement with a sensitive RVP, detected viruses not targeted by the RVP, and yielded epidemiologically and clinically valuable sequence information.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures

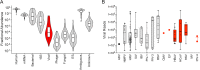

References
-
- Choi SH, Hong SB, Ko GB, Lee Y, Park HJ, Park SY, Moon SM, Cho OH, Park KH, Chong YP, Kim SH, Huh JW, Sung H, Do KH, Lee SO, Kim MN, Jeong JY, Lim CM, Kim YS, Woo JH, Koh Y. 2012. Viral infection in patients with severe pneumonia requiring intensive care unit admission. Am J Respir Crit Care Med 186:325–332. doi: 10.1164/rccm.201112-2240OC. - DOI - PubMed
-
- Jain S, Williams DJ, Arnold SR, Ampofo K, Bramley AM, Reed C, Stockmann C, Anderson EJ, Grijalva CG, Self WH, Zhu Y, Patel A, Hymas W, Chappell JD, Kaufman RA, Kan JH, Dansie D, Lenny N, Hillyard DR, Haynes LM, Levine M, Lindstrom S, Winchell JM, Katz JM, Erdman D, Schneider E, Hicks LA, Wunderink RG, Edwards KM, Pavia AT, McCullers JA, Finelli L, CDC EPIC Study Team. 2015. Community-acquired pneumonia requiring hospitalization among U.S. children. N Engl J Med 372:835–845. doi: 10.1056/NEJMoa1405870. - DOI - PMC - PubMed
-
- Jain S, Self WH, Wunderink RG, Fakhran S, Balk R, Bramley AM, Reed C, Grijalva CG, Anderson EJ, Courtney DM, Chappell JD, Qi C, Hart EM, Carroll F, Trabue C, Donnelly HK, Williams DJ, Zhu Y, Arnold SR, Ampofo K, Waterer GW, Levine M, Lindstrom S, Winchell JM, Katz JM, Erdman D, Schneider E, Hicks LA, McCullers JA, Pavia AT, Edwards KM, Finelli L, CDC EPIC Study Team. 2015. Community-acquired pneumonia requiring hospitalization among U.S. adults. N Engl J Med 373:415–427. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

