T-cell libraries allow simple parallel generation of multiple peptide-specific human T-cell clones
- PMID: 26826277
- PMCID: PMC4783706
- DOI: 10.1016/j.jim.2016.01.014
T-cell libraries allow simple parallel generation of multiple peptide-specific human T-cell clones
Abstract
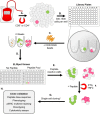
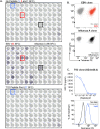
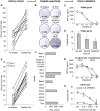
Isolation of peptide-specific T-cell clones is highly desirable for determining the role of T-cells in human disease, as well as for the development of therapies and diagnostics. However, generation of monoclonal T-cells with the required specificity is challenging and time-consuming. Here we describe a library-based strategy for the simple parallel detection and isolation of multiple peptide-specific human T-cell clones from CD8(+) or CD4(+) polyclonal T-cell populations. T-cells were first amplified by CD3/CD28 microbeads in a 96U-well library format, prior to screening for desired peptide recognition. T-cells from peptide-reactive wells were then subjected to cytokine-mediated enrichment followed by single-cell cloning, with the entire process from sample to validated clone taking as little as 6 weeks. Overall, T-cell libraries represent an efficient and relatively rapid tool for the generation of peptide-specific T-cell clones, with applications shown here in infectious disease (Epstein-Barr virus, influenza A, and Ebola virus), autoimmunity (type 1 diabetes) and cancer.
Keywords: Ebola; Library; Peptide-specific; T-cell clone; Tumour; Type 1 diabetes.
Copyright © 2016 The Authors. Published by Elsevier B.V. All rights reserved.
Figures

Similar articles
-
Direct, MHC-dependent presentation of the drug sulfamethoxazole to human alphabeta T cell clones.J Clin Invest. 1997 Jul 1;100(1):136-41. doi: 10.1172/JCI119505. J Clin Invest. 1997. PMID: 9202065 Free PMC article.
-
Evaluation of suitable target antigens and immunoassays for high-accuracy immune monitoring of cytomegalovirus and Epstein-Barr virus-specific T cells as targets of interest in immunotherapeutic approaches.J Immunol Methods. 2014 Jun;408:101-13. doi: 10.1016/j.jim.2014.05.011. Epub 2014 May 28. J Immunol Methods. 2014. PMID: 24877879
-
A novel strategy for rapid and efficient isolation of human tumor-specific CD4(+) and CD8(+) T-cell clones.J Immunol Methods. 2008 Feb 29;331(1-2):13-26. doi: 10.1016/j.jim.2007.09.006. Epub 2007 Oct 9. J Immunol Methods. 2008. PMID: 17959194 Free PMC article.
-
Multiplex and functional detection of antigen-specific human T cells by ITRA--indirect T cell recognition assay.J Immunol Methods. 2014 Feb;404:13-23. doi: 10.1016/j.jim.2013.11.027. Epub 2013 Dec 12. J Immunol Methods. 2014. PMID: 24333463
-
Differences in the regulation of CD4 and CD8 T-cell clones during immune responses.Philos Trans R Soc Lond B Biol Sci. 2000 Mar 29;355(1395):401-6. doi: 10.1098/rstb.2000.0580. Philos Trans R Soc Lond B Biol Sci. 2000. PMID: 10794061 Free PMC article. Review.
Cited by
-
Conserved peptide vaccine candidates containing multiple Ebola nucleoprotein epitopes display interactions with diverse HLA molecules.Med Microbiol Immunol. 2019 Apr;208(2):227-238. doi: 10.1007/s00430-019-00584-y. Epub 2019 Feb 21. Med Microbiol Immunol. 2019. PMID: 30790057
-
Human leukocyte antigen (HLA) class II peptide flanking residues tune the immunogenicity of a human tumor-derived epitope.J Biol Chem. 2019 Dec 27;294(52):20246-20258. doi: 10.1074/jbc.RA119.009437. Epub 2019 Oct 16. J Biol Chem. 2019. PMID: 31619516 Free PMC article.
-
Structural definition of HLA class II-presented SARS-CoV-2 epitopes reveals a mechanism to escape pre-existing CD4+ T cell immunity.Cell Rep. 2023 Aug 29;42(8):112827. doi: 10.1016/j.celrep.2023.112827. Epub 2023 Jul 19. Cell Rep. 2023. PMID: 37471227 Free PMC article.
-
T-cell Receptors Engineered De Novo for Peptide Specificity Can Mediate Optimal T-cell Activity without Self Cross-Reactivity.Cancer Immunol Res. 2019 Dec;7(12):2025-2035. doi: 10.1158/2326-6066.CIR-19-0035. Epub 2019 Sep 23. Cancer Immunol Res. 2019. PMID: 31548259 Free PMC article.
-
Hemophilia A inhibitor treatment: the promise of engineered T-cell therapy.Transl Res. 2017 Sep;187:44-52. doi: 10.1016/j.trsl.2017.06.002. Epub 2017 Jun 9. Transl Res. 2017. PMID: 28651073 Free PMC article. Review.
References
-
- Bednarek M.A., Sauma S.Y., Gammon C.C., Porter G., Tamhankar S., Williamson A.R., Zweerink H.J. The minimum peptide epitope from the influenza virus matrix protein. Extra and intracellular loading of HLA-A2. J. Immunol. 1991;147:4047–4053. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

