Genome-Wide Identification, Characterization and Expression Analysis of the Chalcone Synthase Family in Maize
- PMID: 26828478
- PMCID: PMC4783895
- DOI: 10.3390/ijms17020161
Genome-Wide Identification, Characterization and Expression Analysis of the Chalcone Synthase Family in Maize
Abstract
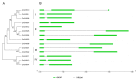
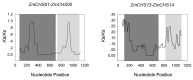
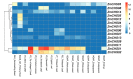
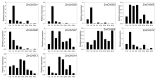
Members of the chalcone synthase (CHS) family participate in the synthesis of a series of secondary metabolites in plants, fungi and bacteria. The metabolites play important roles in protecting land plants against various environmental stresses during the evolutionary process. Our research was conducted on comprehensive investigation of CHS genes in maize (Zea mays L.), including their phylogenetic relationships, gene structures, chromosomal locations and expression analysis. Fourteen CHS genes (ZmCHS01-14) were identified in the genome of maize, representing one of the largest numbers of CHS family members identified in one organism to date. The gene family was classified into four major classes (classes I-IV) based on their phylogenetic relationships. Most of them contained two exons and one intron. The 14 genes were unevenly located on six chromosomes. Two segmental duplication events were identified, which might contribute to the expansion of the maize CHS gene family to some extent. In addition, quantitative real-time PCR and microarray data analyses suggested that ZmCHS genes exhibited various expression patterns, indicating functional diversification of the ZmCHS genes. Our results will contribute to future studies of the complexity of the CHS gene family in maize and provide valuable information for the systematic analysis of the functions of the CHS gene family.
Keywords: chalcone synthase; evolution; expression; genome-wide analysis; maize.
Figures



Similar articles
-
Segmental and tandem chromosome duplications led to divergent evolution of the chalcone synthase gene family in Phalaenopsis orchids.Ann Bot. 2019 Jan 1;123(1):69-77. doi: 10.1093/aob/mcy136. Ann Bot. 2019. PMID: 30113635 Free PMC article.
-
Genome-wide analysis of the chalcone synthase superfamily genes of Physcomitrella patens.Plant Mol Biol. 2010 Feb;72(3):247-63. doi: 10.1007/s11103-009-9565-z. Epub 2009 Oct 31. Plant Mol Biol. 2010. PMID: 19876746
-
Genome-wide identification and characterisation of F-box family in maize.Mol Genet Genomics. 2013 Nov;288(11):559-77. doi: 10.1007/s00438-013-0769-1. Epub 2013 Aug 9. Mol Genet Genomics. 2013. PMID: 23928825
-
Genome-wide identification and phylogenetic analysis of the chalcone synthase gene family in rice.J Plant Res. 2017 Jan;130(1):95-105. doi: 10.1007/s10265-016-0871-7. Epub 2016 Nov 23. J Plant Res. 2017. PMID: 27878652
-
Molecular evolution of the chalcone synthase multigene family in the morning glory genome.Plant Mol Biol. 2000 Jan;42(1):79-92. Plant Mol Biol. 2000. PMID: 10688131 Review.
Cited by
-
Genome-wide identification and expression analysis of the class III peroxidase gene (PRXIII) family in Medicago sativa L. and its function in the abiotic stress response.BMC Plant Biol. 2025 Apr 8;25(1):443. doi: 10.1186/s12870-025-06470-5. BMC Plant Biol. 2025. PMID: 40200136 Free PMC article.
-
Segmental and tandem chromosome duplications led to divergent evolution of the chalcone synthase gene family in Phalaenopsis orchids.Ann Bot. 2019 Jan 1;123(1):69-77. doi: 10.1093/aob/mcy136. Ann Bot. 2019. PMID: 30113635 Free PMC article.
-
Comparative genomic analysis of the PKS genes in five species and expression analysis in upland cotton.PeerJ. 2017 Oct 30;5:e3974. doi: 10.7717/peerj.3974. eCollection 2017. PeerJ. 2017. PMID: 29104824 Free PMC article.
-
Comparative transcriptome analysis of maize (Zea mays L.) seedlings in response to copper stress.Open Life Sci. 2024 Nov 6;19(1):20220953. doi: 10.1515/biol-2022-0953. eCollection 2024. Open Life Sci. 2024. PMID: 39533982 Free PMC article.
-
Genome-wide characterization of monoacylglycerol lipase (MAGL) gene family in soybean and functional analysis of GmMAGLs in storage lipid metabolism and drought resistance.BMC Genomics. 2025 Jul 1;26(1):625. doi: 10.1186/s12864-025-11813-5. BMC Genomics. 2025. PMID: 40597643 Free PMC article.
References
-
- Schröder J. A family of plant-specific polyketide synthases: Facts and predictions. Trends Plant Sci. 1997;2:373–378. doi: 10.1016/S1360-1385(97)87121-X. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

