Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution
- PMID: 26829225
- PMCID: PMC4762201
- DOI: 10.1038/nature16940
Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution
Abstract
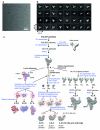
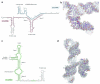
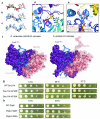
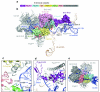
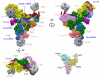
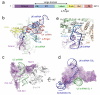
U4/U6.U5 tri-snRNP represents a substantial part of the spliceosome before activation. A cryo-electron microscopy structure of Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at 3.7 Å resolution led to an essentially complete atomic model comprising 30 proteins plus U4/U6 and U5 small nuclear RNAs (snRNAs). The structure reveals striking interweaving interactions of the protein and RNA components, including extended polypeptides penetrating into subunit interfaces. The invariant ACAGAGA sequence of U6 snRNA, which base-pairs with the 5'-splice site during catalytic activation, forms a hairpin stabilized by Dib1 and Prp8 while the adjacent nucleotides interact with the exon binding loop 1 of U5 snRNA. Snu114 harbours GTP, but its putative catalytic histidine is held away from the γ-phosphate by hydrogen bonding to a tyrosine in the amino-terminal domain of Prp8. Mutation of this histidine to alanine has no detectable effect on yeast growth. The structure provides important new insights into the spliceosome activation process leading to the formation of the catalytic centre.
Figures


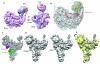



References
-
- Fabrizio P, Dannenberg J, Dube P, Kastner B, Stark H, Urlaub H, Lührmann R. The evolutionarily conserved core design of the catalytic activation step of the yeast spliceosome. Mol. Cell. 2009;36:593–608. - PubMed
-
- Staley JP, Guthrie C. An RNA switch at the 5′ splice requires ATP and the DEAD box protein Prp28p. Mol. Cell. 1999;3:55–64. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

