Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain
- PMID: 26829320
- PMCID: PMC5088052
- DOI: 10.1038/nbt.3440
Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain
Abstract
Recombinant adeno-associated viruses (rAAVs) are commonly used vehicles for in vivo gene transfer. However, the tropism repertoire of naturally occurring AAVs is limited, prompting a search for novel AAV capsids with desired characteristics. Here we describe a capsid selection method, called Cre recombination-based AAV targeted evolution (CREATE), that enables the development of AAV capsids that more efficiently transduce defined Cre-expressing cell populations in vivo. We use CREATE to generate AAV variants that efficiently and widely transduce the adult mouse central nervous system (CNS) after intravenous injection. One variant, AAV-PHP.B, transfers genes throughout the CNS with an efficiency that is at least 40-fold greater than that of the current standard, AAV9 (refs. 14,15,16,17), and transduces the majority of astrocytes and neurons across multiple CNS regions. In vitro, it transduces human neurons and astrocytes more efficiently than does AAV9, demonstrating the potential of CREATE to produce customized AAV vectors for biomedical applications.
Figures

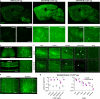
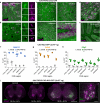

References
-
- Kaplitt MG, et al. Safety and tolerability of gene therapy with an adeno-associated virus (AAV) borne GAD gene for Parkinson's disease: an open label, phase I trial. Lancet. 2007;369:2097–2105. doi:10.1016/S0140-6736(07)60982-9. - PubMed
-
- Wu Z, Asokan A, Samulski RJ. Adeno-associated virus serotypes: vector toolkit for human gene therapy. Molecular therapy : the journal of the American Society of Gene Therapy. 2006;14:316–327. doi:10.1016/j.ymthe.2006.05.009. - PubMed
-
- High KH, Nathwani A, Spencer T, Lillicrap D. Current status of haemophilia gene therapy. Haemophilia : the official journal of the World Federation of Hemophilia 20 Suppl. 2014;4:43–49. doi:10.1111/hae.12411. - PubMed
-
- Ojala DS, Amara DP, Schaffer DV. Adeno-associated virus vectors and neurological gene therapy. Neuroscientist. 2015;21:84–98. doi:10.1177/1073858414521870. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 1R21MH103824-01/MH/NIMH NIH HHS/United States
- DP2 NS087949/NS/NINDS NIH HHS/United States
- 1DP2NS087949/DP/NCCDPHP CDC HHS/United States
- R21 MH103824/MH/NIMH NIH HHS/United States
- 1R01MH100900/MH/NIMH NIH HHS/United States
- R01MH107800/MH/NIMH NIH HHS/United States
- R01 MH107800/MH/NIMH NIH HHS/United States
- U01 NS090577/NS/NINDS NIH HHS/United States
- R01 AG047664/AG/NIA NIH HHS/United States
- R01 MH100900/MH/NIMH NIH HHS/United States
- 1R01AG047664/AG/NIA NIH HHS/United States
- 1U01NS090577/NS/NINDS NIH HHS/United States
- 1R01MH100900-02S1/MH/NIMH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

