Classification of proteins with shared motifs and internal repeats in the ECOD database
- PMID: 26833690
- PMCID: PMC4918410
- DOI: 10.1002/pro.2893
Classification of proteins with shared motifs and internal repeats in the ECOD database
Abstract
Proteins and their domains evolve by a set of events commonly including the duplication and divergence of small motifs. The presence of short repetitive regions in domains has generally constituted a difficult case for structural domain classifications and their hierarchies. We developed the Evolutionary Classification Of protein Domains (ECOD) in part to implement a new schema for the classification of these types of proteins. Here we document the ways in which ECOD classifies proteins with small internal repeats, widespread functional motifs, and assemblies of small domain-like fragments in its evolutionary schema. We illustrate the ways in which the structural genomics project impacted the classification and characterization of new structural domains and sequence families over the decade.
Keywords: internal; protein classification; protein motifs; repeats; structural bioinformatics; structural genomics.
© 2016 The Protein Society.
Figures
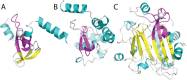




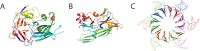



Similar articles
-
ECOD: new developments in the evolutionary classification of domains.Nucleic Acids Res. 2017 Jan 4;45(D1):D296-D302. doi: 10.1093/nar/gkw1137. Epub 2016 Nov 29. Nucleic Acids Res. 2017. PMID: 27899594 Free PMC article.
-
ECOD: an evolutionary classification of protein domains.PLoS Comput Biol. 2014 Dec 4;10(12):e1003926. doi: 10.1371/journal.pcbi.1003926. eCollection 2014 Dec. PLoS Comput Biol. 2014. PMID: 25474468 Free PMC article.
-
ECOD: integrating classifications of protein domains from experimental and predicted structures.Nucleic Acids Res. 2025 Jan 6;53(D1):D411-D418. doi: 10.1093/nar/gkae1029. Nucleic Acids Res. 2025. PMID: 39565196 Free PMC article.
-
Comparison of ARM and HEAT protein repeats.J Mol Biol. 2001 May 25;309(1):1-18. doi: 10.1006/jmbi.2001.4624. J Mol Biol. 2001. PMID: 11491282 Review.
-
Protein repeats: structures, functions, and evolution.J Struct Biol. 2001 May-Jun;134(2-3):117-31. doi: 10.1006/jsbi.2001.4392. J Struct Biol. 2001. PMID: 11551174 Review.
Cited by
-
Influence of circular permutations on the structure and stability of a six-fold circular symmetric designer protein.Protein Sci. 2020 Dec;29(12):2375-2386. doi: 10.1002/pro.3961. Epub 2020 Oct 16. Protein Sci. 2020. PMID: 33006397 Free PMC article.
-
DomainMapper: Accurate domain structure annotation including those with non-contiguous topologies.Protein Sci. 2022 Nov;31(11):e4465. doi: 10.1002/pro.4465. Protein Sci. 2022. PMID: 36208126 Free PMC article.
-
ECOD: identification of distant homology among multidomain and transmembrane domain proteins.BMC Mol Cell Biol. 2019 Jun 21;20(1):18. doi: 10.1186/s12860-019-0204-5. BMC Mol Cell Biol. 2019. PMID: 31226926 Free PMC article.
-
Piecing Together the History of Protein Folds From a Fragmented Evolutionary Record.Genome Biol Evol. 2025 Jul 30;17(8):evaf148. doi: 10.1093/gbe/evaf148. Genome Biol Evol. 2025. PMID: 40839423 Free PMC article.
-
ProLego: tool for extracting and visualizing topological modules in protein structures.BMC Bioinformatics. 2018 May 4;19(1):167. doi: 10.1186/s12859-018-2171-9. BMC Bioinformatics. 2018. PMID: 29728050 Free PMC article.
References
-
- Grishin NV (2001) Fold change in evolution of protein structures. J Struct Biol 134:167–185. - PubMed
-
- Lupas AN, Ponting CP, Russell RB (2001) On the evolution of protein folds: are similar motifs in different protein folds the result of convergence, insertion, or relics of an ancient peptide world? J Struct Biol 134:191–203. - PubMed
-
- Kinch LN, Grishin NV (2002) Evolution of protein structures and functions. Curr Opin Struct Biol 12:400–408. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

