Establishment and use of new MDCK II cells overexpressing both UGT1A1 and MRP2 to characterize flavonoid metabolism via the glucuronidation pathway
- PMID: 26833852
- PMCID: PMC5358013
- DOI: 10.1002/mnfr.201500321
Establishment and use of new MDCK II cells overexpressing both UGT1A1 and MRP2 to characterize flavonoid metabolism via the glucuronidation pathway
Abstract
Scope: The purpose of this study is to characterize how overexpression of an efflux transporter and an UDP-glucuronosyltransferase (UGT) affects the cellular kinetics of glucuronidation processes.
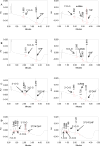
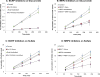
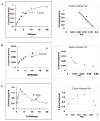
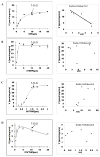
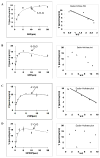
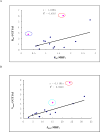
Methods and results: A new MDCK II cell line overexpressing both MRP2 and UGT1A1 (MDCKII-UGT1A1/MRP2 cells) was developed and used to determine how overexpression of an efflux transporter affects the kinetics of cellular flavonoid glucuronide production. The results showed that most model flavonoids (from a total of 13) were mainly metabolized into glucuronides in the MDCKII-UGT1A1/MRP2 cells and the glucuronides were rapidly excreted. Flavonoids with three or fewer hydroxyl group at 7, 3' or 6 hydroxyl group were also metabolized into sulfates. Mechanistic studies using 7-hydroxylflavone showed that its glucuronide was mainly (90%) effluxed by BCRP with a small (10%) but significant contribution from MRP2. Maximal velocity of glucuronide production MDCK-MRP2/UGT1A1 cells showed a fairly good correlation (R(2) >0.8) with those derived using UGT1A1 microsomes, but other kinetic parameters (e.g., Km ) did not correlate.
Conclusion: Overexpression of a second efficient efflux transporter did not significantly change the fact that BCRP is the dominant transporter for flavonoid glucuronide nor did it diminish the influence of the efflux transporter as the "gate keeper" of glucuronidation process.
Keywords: Efflux transporters; Flavonoids; Glucuronide; UGT; “Revolving door”.
© 2016 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures


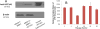







Similar articles
-
Glucuronidation: driving factors and their impact on glucuronide disposition.Drug Metab Rev. 2017 May;49(2):105-138. doi: 10.1080/03602532.2017.1293682. Epub 2017 May 22. Drug Metab Rev. 2017. PMID: 28266877 Free PMC article. Review.
-
Chemical inhibition and stable knock-down of efflux transporters leads to reduced glucuronidation of wushanicaritin in UGT1A1-overexpressing HeLa cells: the role of breast cancer resistance protein (BCRP) and multidrug resistance-associated proteins (MRPs) in the excretion of glucuronides.Food Funct. 2018 Mar 1;9(3):1410-1423. doi: 10.1039/c7fo01298e. Epub 2018 Jan 10. Food Funct. 2018. PMID: 29318243
-
UDP-glucuronosyltransferase (UGT) 1A9-overexpressing HeLa cells is an appropriate tool to delineate the kinetic interplay between breast cancer resistance protein (BRCP) and UGT and to rapidly identify the glucuronide substrates of BCRP.Drug Metab Dispos. 2012 Feb;40(2):336-45. doi: 10.1124/dmd.111.041467. Epub 2011 Nov 9. Drug Metab Dispos. 2012. PMID: 22071170 Free PMC article.
-
Curcumin Affects Phase II Disposition of Resveratrol Through Inhibiting Efflux Transporters MRP2 and BCRP.Pharm Res. 2016 Mar;33(3):590-602. doi: 10.1007/s11095-015-1812-1. Epub 2015 Oct 26. Pharm Res. 2016. PMID: 26502886 Free PMC article.
-
An update on polyphenol disposition via coupled metabolic pathways.Expert Opin Drug Metab Toxicol. 2019 Feb;15(2):151-165. doi: 10.1080/17425255.2019.1559815. Epub 2018 Dec 24. Expert Opin Drug Metab Toxicol. 2019. PMID: 30583703 Review.
Cited by
-
Glucuronidation: driving factors and their impact on glucuronide disposition.Drug Metab Rev. 2017 May;49(2):105-138. doi: 10.1080/03602532.2017.1293682. Epub 2017 May 22. Drug Metab Rev. 2017. PMID: 28266877 Free PMC article. Review.
-
Potential metabolism determinants and drug-drug interactions of a natural flavanone bavachinin.RSC Adv. 2020 Sep 23;10(58):35141-35152. doi: 10.1039/d0ra06961b. eCollection 2020 Sep 21. RSC Adv. 2020. PMID: 35515695 Free PMC article.
-
The Efflux Mechanism of Fraxetin-O-Glucuronides in UGT1A9-Transfected HeLa Cells: Identification of Multidrug Resistance-Associated Proteins 3 and 4 (MRP3/4) as the Important Contributors.Front Pharmacol. 2019 May 7;10:496. doi: 10.3389/fphar.2019.00496. eCollection 2019. Front Pharmacol. 2019. PMID: 31133859 Free PMC article.
-
Mechanism of the efflux transport of demethoxycurcumin-O-glucuronides in HeLa cells stably transfected with UDP-glucuronosyltransferase 1A1.PLoS One. 2019 May 31;14(5):e0217695. doi: 10.1371/journal.pone.0217695. eCollection 2019. PLoS One. 2019. PMID: 31150474 Free PMC article.
-
The roles of breast cancer resistance protein (BCRP/ABCG2) and multidrug resistance-associated proteins (MRPs/ABCCs) in the excretion of cycloicaritin-3-O-glucoronide in UGT1A1-overexpressing HeLa cells.Chem Biol Interact. 2018 Dec 25;296:45-56. doi: 10.1016/j.cbi.2018.09.001. Epub 2018 Sep 17. Chem Biol Interact. 2018. PMID: 30237061 Free PMC article.
References
-
- Fedejko B, Mazerska Z. UDP-glucuronyltransferases in detoxification and activation metabolism of endogenous compounds and xenobiotics. Postepy Biochem. 2011;57:49–62. - PubMed
-
- Radominska-Pandya A, Bratton S, Little JM. A historical overview of the heterologous expression of mammalian UDP-glucuronosyltransferase isoforms over the past twenty years. Curr Drug Metab. 2005;6:141–160. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

