RAG-mediated DNA double-strand breaks activate a cell type-specific checkpoint to inhibit pre-B cell receptor signals
- PMID: 26834154
- PMCID: PMC4749927
- DOI: 10.1084/jem.20151048
RAG-mediated DNA double-strand breaks activate a cell type-specific checkpoint to inhibit pre-B cell receptor signals
Abstract
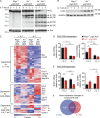
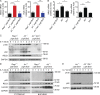
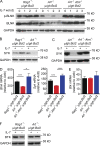
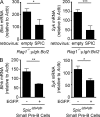
DNA double-strand breaks (DSBs) activate a canonical DNA damage response, including highly conserved cell cycle checkpoint pathways that prevent cells with DSBs from progressing through the cell cycle. In developing B cells, pre-B cell receptor (pre-BCR) signals initiate immunoglobulin light (Igl) chain gene assembly, leading to RAG-mediated DNA DSBs. The pre-BCR also promotes cell cycle entry, which could cause aberrant DSB repair and genome instability in pre-B cells. Here, we show that RAG DSBs inhibit pre-BCR signals through the ATM- and NF-κB2-dependent induction of SPIC, a hematopoietic-specific transcriptional repressor. SPIC inhibits expression of the SYK tyrosine kinase and BLNK adaptor, resulting in suppression of pre-BCR signaling. This regulatory circuit prevents the pre-BCR from inducing additional Igl chain gene rearrangements and driving pre-B cells with RAG DSBs into cycle. We propose that pre-B cells toggle between pre-BCR signals and a RAG DSB-dependent checkpoint to maintain genome stability while iteratively assembling Igl chain genes.
© 2016 Bednarski et al.
Figures



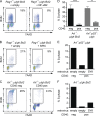
References
-
- Bredemeyer A.L., Helmink B.A., Innes C.L., Calderon B., McGinnis L.M., Mahowald G.K., Gapud E.J., Walker L.M., Collins J.B., Weaver B.K., et al. 2008. DNA double-strand breaks activate a multi-functional genetic program in developing lymphocytes. Nature. 456:819–823. 10.1038/nature07392 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- T32 CA113275/CA/NCI NIH HHS/United States
- R01 AI112621/AI/NIAID NIH HHS/United States
- K08AI106953/AI/NIAID NIH HHS/United States
- K08 AI102946/AI/NIAID NIH HHS/United States
- AI47829/AI/NIAID NIH HHS/United States
- R01AI112621/AI/NIAID NIH HHS/United States
- T32 HD007499/HD/NICHD NIH HHS/United States
- K08AI102946-01/AI/NIAID NIH HHS/United States
- K08 AI106953/AI/NIAID NIH HHS/United States
- P30 CA091842/CA/NCI NIH HHS/United States
- UL1 TR000448/TR/NCATS NIH HHS/United States
- R01 AI074953/AI/NIAID NIH HHS/United States
- R01 CA136470/CA/NCI NIH HHS/United States
- CA136470/CA/NCI NIH HHS/United States
- AI074953/AI/NIAID NIH HHS/United States
- R01 AI047829/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

