Metagenomic analysis of microbial consortia enriched from compost: new insights into the role of Actinobacteria in lignocellulose decomposition
- PMID: 26834834
- PMCID: PMC4731972
- DOI: 10.1186/s13068-016-0440-2
Metagenomic analysis of microbial consortia enriched from compost: new insights into the role of Actinobacteria in lignocellulose decomposition
Abstract
Background: Compost habitats sustain a vast ensemble of microbes specializing in the degradation of lignocellulosic plant materials and are thus important both for their roles in the global carbon cycle and as potential sources of biochemical catalysts for advanced biofuels production. Studies have revealed substantial diversity in compost microbiomes, yet how this diversity relates to functions and even to the genes encoding lignocellulolytic enzymes remains obscure. Here, we used a metagenomic analysis of the rice straw-adapted (RSA) microbial consortia enriched from compost ecosystems to decipher the systematic and functional contexts within such a distinctive microbiome.
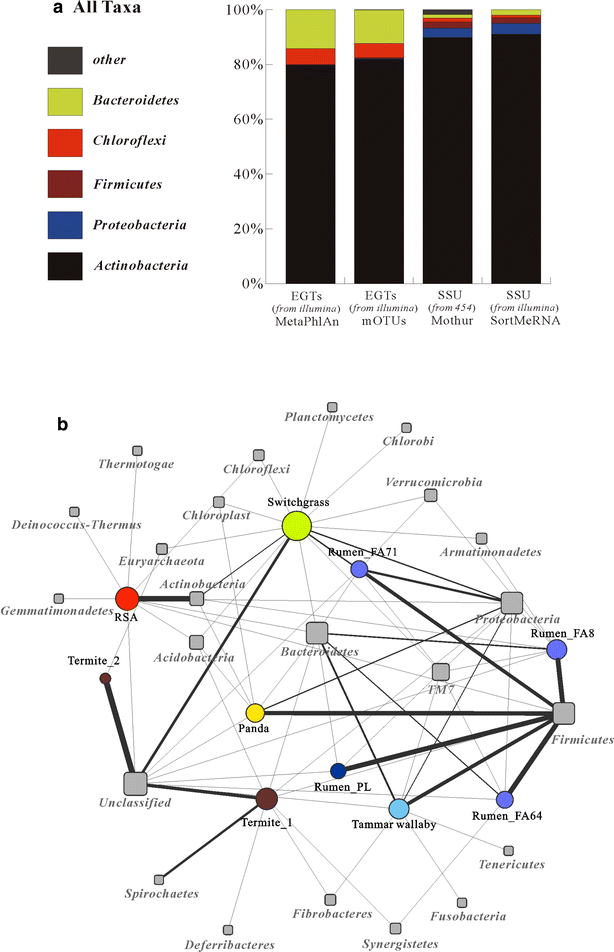
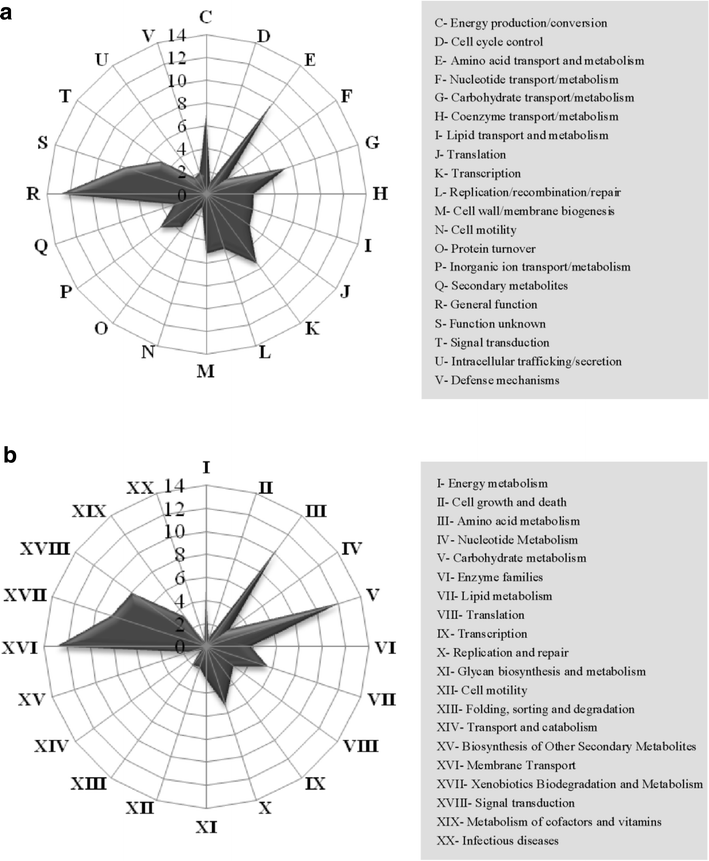
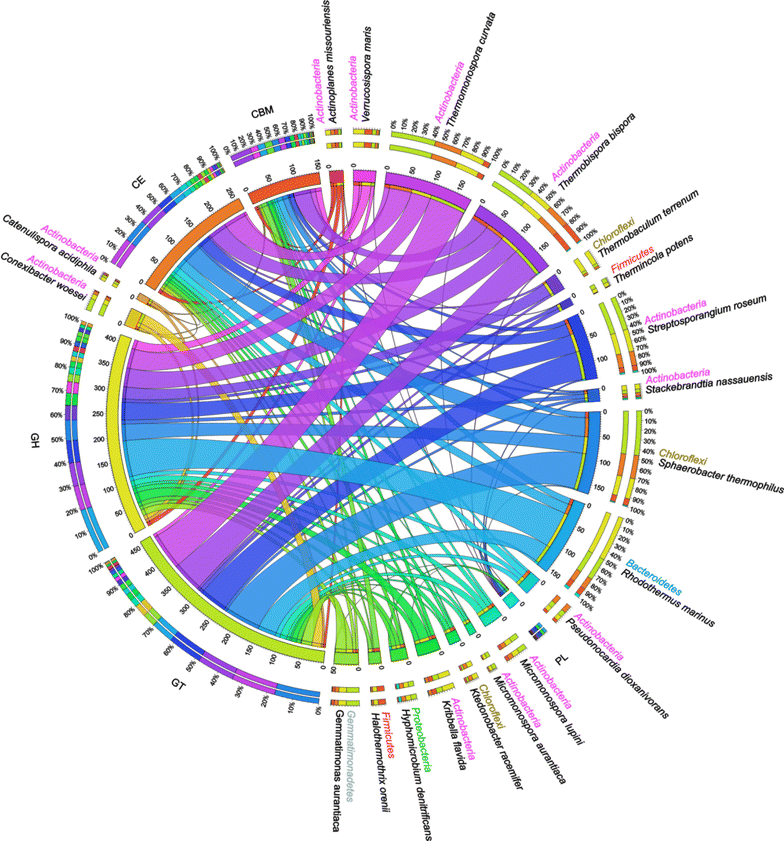
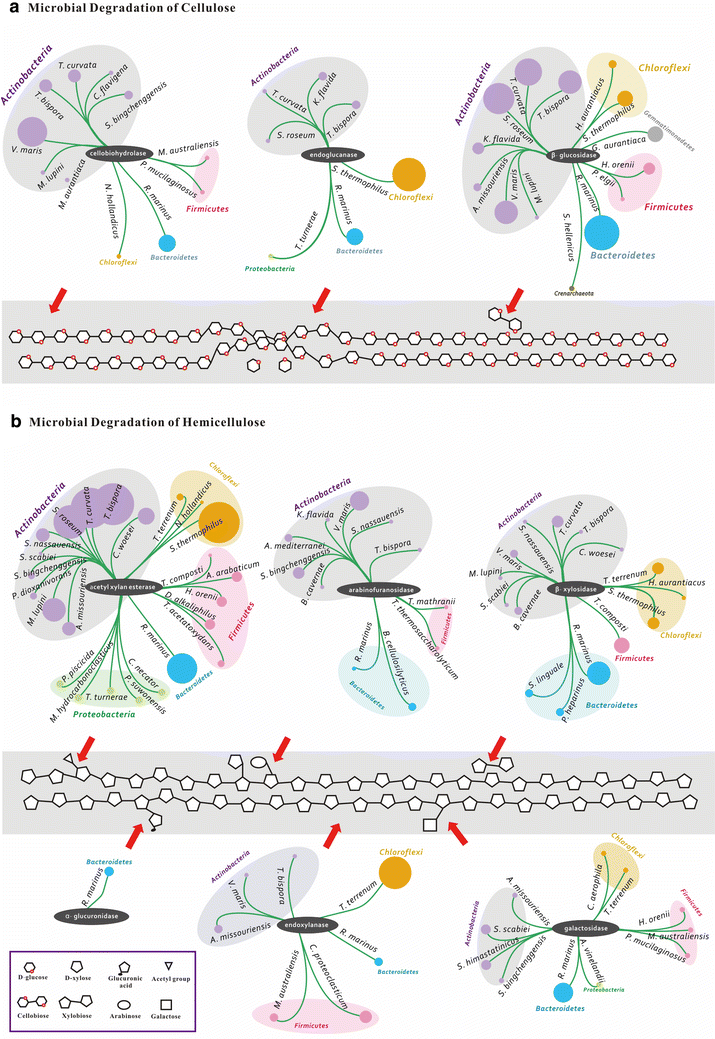
Results: Analyses of the 16S pyrotag library and 5 Gbp of metagenomic sequence showed that the phylum Actinobacteria was the predominant group among the Bacteria in the RSA consortia, followed by Proteobacteria, Firmicutes, Chloroflexi, and Bacteroidetes. The CAZymes profile revealed that CAZyme genes in the RSA consortia were also widely distributed within these bacterial phyla. Strikingly, about 46.1 % of CAZyme genes were from actinomycetal communities, which harbored a substantially expanded catalog of the cellobiohydrolase, β-glucosidase, acetyl xylan esterase, arabinofuranosidase, pectin lyase, and ligninase genes. Among these communities, a variety of previously unrecognized species was found, which reveals a greater ecological functional diversity of thermophilic Actinobacteria than previously assumed.
Conclusion: These data underline the pivotal role of thermophilic Actinobacteria in lignocellulose biodegradation processes in the compost habitat. Besides revealing a new benchmark for microbial enzymatic deconstruction of lignocelluloses, the results suggest that actinomycetes found in compost ecosystems are potential candidates for mining efficient lignocellulosic enzymes in the biofuel industry.
Keywords: Actinobacteria; Compost ecosystem; Lignocellulose degradation; Metagenomics.
Figures
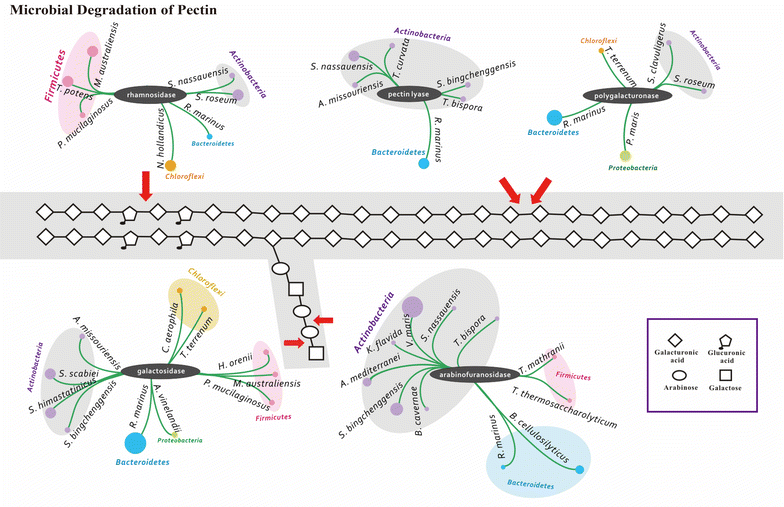
References
-
- Kumar P, Barrett DM, Delwiche MJ, Stroeve P. Methods for pretreatment of lignocellulosic biomass for efficient hydrolysis and biofuel production. Ind Eng Chem Res. 2009;48(8):3713–3729. doi: 10.1021/ie801542g. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

