Dynamic DNA binding licenses a repair factor to bypass roadblocks in search of DNA lesions
- PMID: 26837705
- PMCID: PMC4742970
- DOI: 10.1038/ncomms10607
Dynamic DNA binding licenses a repair factor to bypass roadblocks in search of DNA lesions
Abstract
DNA-binding proteins search for specific targets via facilitated diffusion along a crowded genome. However, little is known about how crowded DNA modulates facilitated diffusion and target recognition. Here we use DNA curtains and single-molecule fluorescence imaging to investigate how Msh2-Msh3, a eukaryotic mismatch repair complex, navigates on crowded DNA. Msh2-Msh3 hops over nucleosomes and other protein roadblocks, but maintains sufficient contact with DNA to recognize a single lesion. In contrast, Msh2-Msh6 slides without hopping and is largely blocked by protein roadblocks. Remarkably, the Msh3-specific mispair-binding domain (MBD) licences a chimeric Msh2-Msh6(3MBD) to bypass nucleosomes. Our studies contrast how Msh2-Msh3 and Msh2-Msh6 navigate on a crowded genome and suggest how Msh2-Msh3 locates DNA lesions outside of replication-coupled repair. These results also provide insights into how DNA repair factors search for DNA lesions in the context of chromatin.
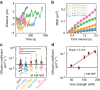
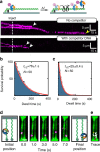
Figures




Similar articles
-
Mispair-specific recruitment of the Mlh1-Pms1 complex identifies repair substrates of the Saccharomyces cerevisiae Msh2-Msh3 complex.J Biol Chem. 2014 Mar 28;289(13):9352-64. doi: 10.1074/jbc.M114.552190. Epub 2014 Feb 18. J Biol Chem. 2014. PMID: 24550389 Free PMC article.
-
Saccharomyces cerevisiae MSH2-MSH3 and MSH2-MSH6 complexes display distinct requirements for DNA binding domain I in mismatch recognition.J Mol Biol. 2007 Feb 9;366(1):53-66. doi: 10.1016/j.jmb.2006.10.099. Epub 2006 Nov 3. J Mol Biol. 2007. PMID: 17157869 Free PMC article.
-
Prerecognition Diffusion Mechanism of Human DNA Mismatch Repair Proteins along DNA: Msh2-Msh3 versus Msh2-Msh6.Biochemistry. 2020 Dec 29;59(51):4822-4832. doi: 10.1021/acs.biochem.0c00669. Epub 2020 Dec 15. Biochemistry. 2020. PMID: 33319999 Free PMC article.
-
DNA binding properties of the yeast Msh2-Msh6 and Mlh1-Pms1 heterodimers.Biol Chem. 2002 Jun;383(6):969-75. doi: 10.1515/BC.2002.103. Biol Chem. 2002. PMID: 12222686 Review.
-
Elucidating the role of interacting residues of the MSH2-MSH6 complex in DNA repair mechanism: A computational approach.Adv Protein Chem Struct Biol. 2019;115:325-350. doi: 10.1016/bs.apcsb.2018.11.005. Epub 2019 Jan 7. Adv Protein Chem Struct Biol. 2019. PMID: 30798936 Review.
Cited by
-
Single-molecule fluorescence imaging techniques reveal molecular mechanisms underlying deoxyribonucleic acid damage repair.Front Bioeng Biotechnol. 2022 Sep 15;10:973314. doi: 10.3389/fbioe.2022.973314. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36185427 Free PMC article. Review.
-
Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair.Mol Cell. 2017 Sep 7;67(5):891-898.e4. doi: 10.1016/j.molcel.2017.08.002. Epub 2017 Aug 31. Mol Cell. 2017. PMID: 28867292 Free PMC article.
-
The crosstalk between telomeres and DNA repair mechanisms: an overview to mammalian somatic cells, germ cells, and preimplantation embryos.J Assist Reprod Genet. 2024 Feb;41(2):277-291. doi: 10.1007/s10815-023-03008-2. Epub 2024 Jan 2. J Assist Reprod Genet. 2024. PMID: 38165506 Free PMC article. Review.
-
An Msh3 ATPase domain mutation has no effect on MMR function.BMC Res Notes. 2017 Nov 25;10(1):616. doi: 10.1186/s13104-017-2939-4. BMC Res Notes. 2017. PMID: 29178930 Free PMC article.
-
ATP binding facilitates target search of SWR1 chromatin remodeler by promoting one-dimensional diffusion on DNA.Elife. 2022 Jul 25;11:e77352. doi: 10.7554/eLife.77352. Elife. 2022. PMID: 35876491 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

