Co-expression of active human cytochrome P450 1A2 and cytochrome P450 reductase on the cell surface of Escherichia coli
- PMID: 26838175
- PMCID: PMC4736170
- DOI: 10.1186/s12934-016-0427-5
Co-expression of active human cytochrome P450 1A2 and cytochrome P450 reductase on the cell surface of Escherichia coli
Abstract
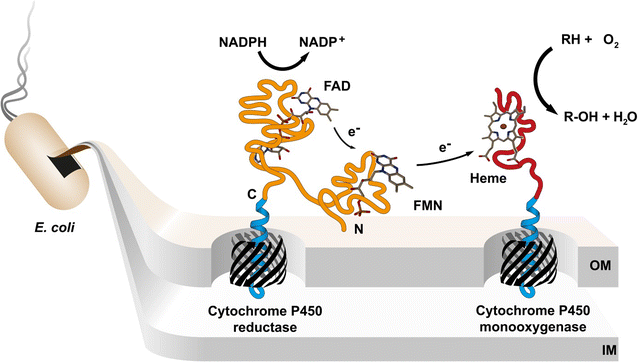
Background: Human cytochrome P450 (CYP) enzymes mediate the first step in the breakdown of most drugs and are strongly involved in drug-drug interactions, drug clearance and activation of prodrugs. Their biocatalytic behavior is a key parameter during drug development which requires preparative synthesis of CYP related drug metabolites. However, recombinant expression of CYP enzymes is a challenging bottleneck for drug metabolite biosynthesis. Therefore, we developed a novel approach by displaying human cytochrome P450 1A2 (CYP1A2) and cytochrome P450 reductase (CPR) on the surface of Escherichia coli.
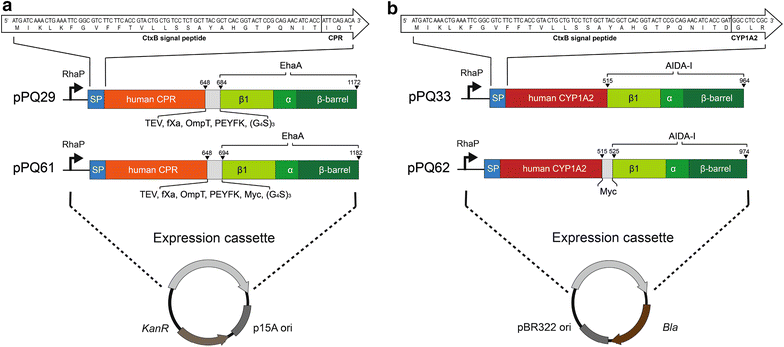
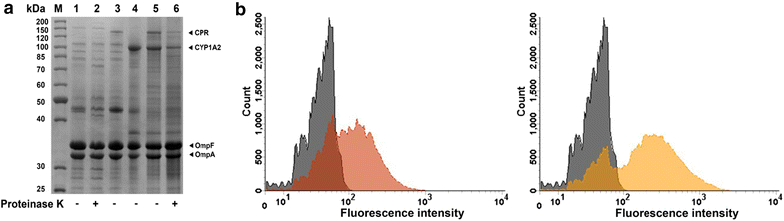
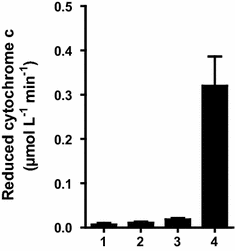
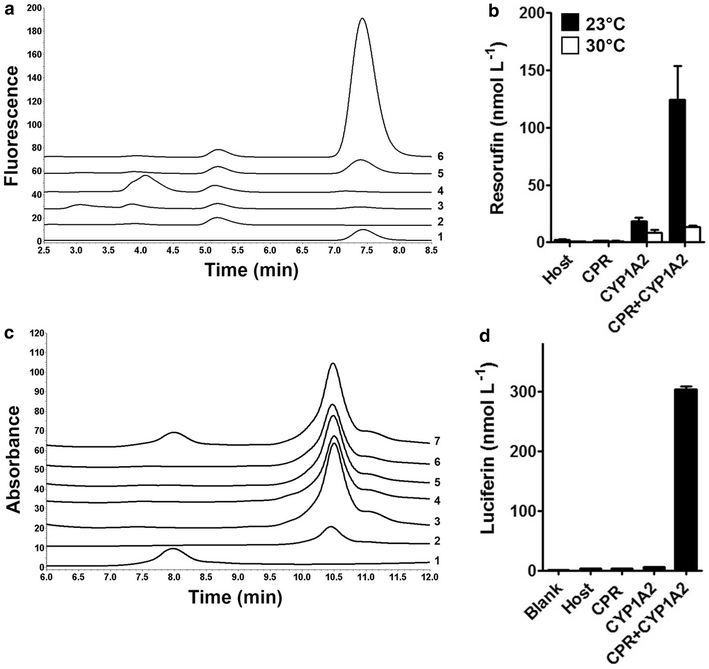
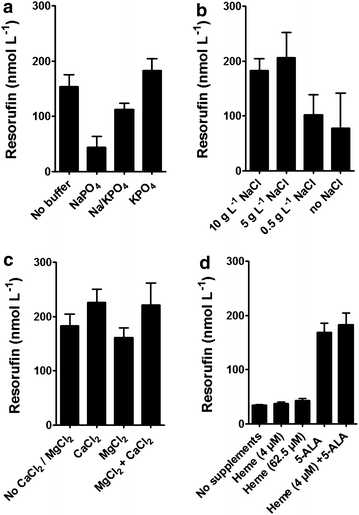
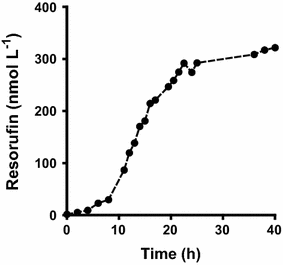
Results: To present human CYP1A2 and CPR on the surface, we employed autodisplay. Both enzymes were displayed on the surface which was demonstrated by protease and antibody accessibility tests. CPR activity was first confirmed with the protein substrate cytochrome c. Cells co-expressing CYP1A2 and CPR were capable of catalyzing the conversion of the known CYP1A2 substrates 7-ethoxyresorufin, phenacetin and the artificial substrate luciferin-MultiCYP, which would not have been possible without interaction of both enzymes. Biocatalytic activity was strongly influenced by the composition of the growth medium. Addition of 5-aminolevulinic acid was necessary to obtain a fully active whole cell biocatalyst and was superior to the addition of heme.
Conclusion: We demonstrated that CYP1A2 and CPR can be co-expressed catalytically active on the cell surface of E. coli. It is a promising step towards pharmaceutical applications such as the synthesis of drug metabolites.
Figures
Similar articles
-
Improving the activity of surface displayed cytochrome P450 enzymes by optimizing the outer membrane linker.Biochim Biophys Acta Biomembr. 2017 Jan;1859(1):104-116. doi: 10.1016/j.bbamem.2016.10.022. Epub 2016 Nov 1. Biochim Biophys Acta Biomembr. 2017. PMID: 27814979
-
Increased Phenacetin Oxidation upon the L382V Substitution in Cytochrome P450 1A2 is Associated with Altered Substrate Binding Orientation.Int J Mol Sci. 2018 May 25;19(6):1580. doi: 10.3390/ijms19061580. Int J Mol Sci. 2018. PMID: 29799514 Free PMC article.
-
Preparative synthesis of drug metabolites using human cytochrome P450s 3A4, 2C9 and 1A2 with NADPH-P450 reductase expressed in Escherichia coli.J Ind Microbiol Biotechnol. 2005 Feb;32(2):67-74. doi: 10.1007/s10295-004-0202-1. Epub 2005 Mar 1. J Ind Microbiol Biotechnol. 2005. PMID: 15739102
-
Metabolism and Mechanism of Human Cytochrome P450 Enzyme 1A2.Curr Drug Metab. 2021;22(1):40-49. doi: 10.2174/1389200221999210101233135. Curr Drug Metab. 2021. PMID: 33397254 Review.
-
Mechanisms of cytochrome P450 1A2-mediated formation of N-hydroxy arylamines and heterocyclic amines and their reaction with guanyl residues.Princess Takamatsu Symp. 1995;23:78-84. Princess Takamatsu Symp. 1995. PMID: 8844798 Review.
Cited by
-
Layer-by-Layer Assembly of Nanosized Membrane Fractions for the Assessment of Cytochrome P450 Xenobiotic Metabolism.ACS Omega. 2018 Oct 3;3(10):12535-12544. doi: 10.1021/acsomega.8b01738. eCollection 2018 Oct 31. ACS Omega. 2018. PMID: 31457987 Free PMC article.
-
Precise Engineering and Efficient Biosynthesis of Robust and High-Activity Human Haemoglobin for Artificial Oxygen Carriers.Microb Biotechnol. 2025 Mar;18(3):e70128. doi: 10.1111/1751-7915.70128. Microb Biotechnol. 2025. PMID: 40072822 Free PMC article.
-
Whole-Cell P450 Biocatalysis Using Engineered Escherichia coli with Fine-Tuned Heme Biosynthesis.Adv Sci (Weinh). 2023 Feb;10(6):e2205580. doi: 10.1002/advs.202205580. Epub 2022 Dec 16. Adv Sci (Weinh). 2023. PMID: 36526588 Free PMC article.
-
Comparison of human cytochrome P450 1A1-catalysed oxidation of benzo[a]pyrene in prokaryotic and eukaryotic expression systems.Monatsh Chem. 2017;148(11):1959-1969. doi: 10.1007/s00706-017-2002-0. Epub 2017 Jul 10. Monatsh Chem. 2017. PMID: 29104317 Free PMC article.
-
Combining lipid-mimicking-enabled transition metal and enzyme-mediated catalysis at the cell surface of E. coli.Chem Sci. 2023 Oct 6;14(42):11896-11906. doi: 10.1039/d3sc02960c. eCollection 2023 Nov 1. Chem Sci. 2023. PMID: 37920346 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

