Early Folding Events, Local Interactions, and Conservation of Protein Backbone Rigidity
- PMID: 26840723
- PMCID: PMC4744172
- DOI: 10.1016/j.bpj.2015.12.028
Early Folding Events, Local Interactions, and Conservation of Protein Backbone Rigidity
Abstract
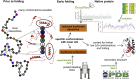
Protein folding is in its early stages largely determined by the protein sequence and complex local interactions between amino acids, resulting in lower energy conformations that provide the context for further folding into the native state. We compiled a comprehensive data set of early folding residues based on pulsed labeling hydrogen deuterium exchange experiments. These early folding residues have corresponding higher backbone rigidity as predicted by DynaMine from sequence, an effect also present when accounting for the secondary structures in the folded protein. We then show that the amino acids involved in early folding events are not more conserved than others, but rather, early folding fragments and the secondary structure elements they are part of show a clear trend toward conserving a rigid backbone. We therefore propose that backbone rigidity is a fundamental physical feature conserved by proteins that can provide important insights into their folding mechanisms and stability.
Copyright © 2016 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures




Similar articles
-
Structure of a hydrophobically collapsed intermediate on the conformational folding pathway of ribonuclease A probed by hydrogen-deuterium exchange.Biochemistry. 1996 Sep 10;35(36):11734-46. doi: 10.1021/bi961085c. Biochemistry. 1996. PMID: 8794754
-
Exploring the Sequence-based Prediction of Folding Initiation Sites in Proteins.Sci Rep. 2017 Aug 18;7(1):8826. doi: 10.1038/s41598-017-08366-3. Sci Rep. 2017. PMID: 28821744 Free PMC article.
-
Folding stability and cooperativity of the three forms of 1-110 residues fragment of staphylococcal nuclease.Biophys J. 2007 Mar 15;92(6):2090-107. doi: 10.1529/biophysj.106.092155. Epub 2006 Dec 15. Biophys J. 2007. PMID: 17172296 Free PMC article.
-
Cytochrome c folding dynamics.Curr Opin Chem Biol. 2004 Apr;8(2):169-74. doi: 10.1016/j.cbpa.2004.02.009. Curr Opin Chem Biol. 2004. PMID: 15062778 Review.
-
The hydrogen exchange core and protein folding.Protein Sci. 1999 Aug;8(8):1571-90. doi: 10.1110/ps.8.8.1571. Protein Sci. 1999. PMID: 10452602 Free PMC article. Review.
Cited by
-
Characterizing the relation of functional and Early Folding Residues in protein structures using the example of aminoacyl-tRNA synthetases.PLoS One. 2018 Oct 30;13(10):e0206369. doi: 10.1371/journal.pone.0206369. eCollection 2018. PLoS One. 2018. PMID: 30376559 Free PMC article.
-
Auto-encoding NMR chemical shifts from their native vector space to a residue-level biophysical index.Nat Commun. 2019 Jun 7;10(1):2511. doi: 10.1038/s41467-019-10322-w. Nat Commun. 2019. PMID: 31175284 Free PMC article.
-
Energy Bilocalization Effect and the Emergence of Molecular Functions in Proteins.Front Mol Biosci. 2021 Dec 23;8:736376. doi: 10.3389/fmolb.2021.736376. eCollection 2021. Front Mol Biosci. 2021. PMID: 35004841 Free PMC article.
-
Mapping the Geometric Evolution of Protein Folding Motor.PLoS One. 2016 Oct 7;11(10):e0163993. doi: 10.1371/journal.pone.0163993. eCollection 2016. PLoS One. 2016. PMID: 27716851 Free PMC article.
-
Observation selection bias in contact prediction and its implications for structural bioinformatics.Sci Rep. 2016 Nov 18;6:36679. doi: 10.1038/srep36679. Sci Rep. 2016. PMID: 27857150 Free PMC article.
References
-
- Bajaj M., Blundell T. Evolution and the tertiary structure of proteins. Annu. Rev. Biophys. Bioeng. 1984;13:453–492. - PubMed
-
- Luheshi L.M., Crowther D.C., Dobson C.M. Protein misfolding and disease: from the test tube to the organism. Curr. Opin. Chem. Biol. 2008;12:25–31. - PubMed
-
- Süel G.M., Lockless S.W., Ranganathan R. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nat. Struct. Biol. 2003;10:59–69. - PubMed
-
- Mirny L., Shakhnovich E. Evolutionary conservation of the folding nucleus. J. Mol. Biol. 2001;308:123–129. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

