Disentangling Random Motion and Flow in a Complex Medium
- PMID: 26840734
- PMCID: PMC4744162
- DOI: 10.1016/j.bpj.2015.11.008
Disentangling Random Motion and Flow in a Complex Medium
Erratum in
-
Disentangling Random Motion and Flow in a Complex Medium.Biophys J. 2017 Sep 5;113(5):1173-1174. doi: 10.1016/j.bpj.2017.08.016. Biophys J. 2017. PMID: 28877498 Free PMC article. No abstract available.
Abstract
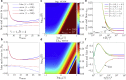
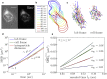
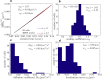
We describe a technique for deconvolving the stochastic motion of particles from large-scale fluid flow in a dynamic environment such as that found in living cells. The method leverages the separation of timescales to subtract out the persistent component of motion from single-particle trajectories. The mean-squared displacement of the resulting trajectories is rescaled so as to enable robust extraction of the diffusion coefficient and subdiffusive scaling exponent of the stochastic motion. We demonstrate the applicability of the method for characterizing both diffusive and fractional Brownian motion overlaid by flow and analytically calculate the accuracy of the method in different parameter regimes. This technique is employed to analyze the motion of lysosomes in motile neutrophil-like cells, showing that the cytoplasm of these cells behaves as a viscous fluid at the timescales examined.
Copyright © 2016 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Intracellular microrheology of motile Amoeba proteus.Biophys J. 2008 Apr 15;94(8):3313-22. doi: 10.1529/biophysj.107.123851. Epub 2008 Jan 11. Biophys J. 2008. PMID: 18192370 Free PMC article.
-
Subdiffusive Dynamics Lead to Depleted Particle Densities near Cellular Borders.Biophys J. 2019 Apr 23;116(8):1538-1546. doi: 10.1016/j.bpj.2019.02.021. Epub 2019 Feb 28. Biophys J. 2019. PMID: 30954212 Free PMC article.
-
Meaningful interpretation of subdiffusive measurements in living cells (crowded environment) by fluorescence fluctuation microscopy.Curr Pharm Biotechnol. 2010 Aug;11(5):527-43. doi: 10.2174/138920110791591454. Curr Pharm Biotechnol. 2010. PMID: 20553227 Free PMC article.
-
Technological strategies to estimate and control diffusive passage times through the mucus barrier in mucosal drug delivery.Adv Drug Deliv Rev. 2018 Jan 15;124:64-81. doi: 10.1016/j.addr.2017.12.002. Epub 2017 Dec 12. Adv Drug Deliv Rev. 2018. PMID: 29246855 Free PMC article. Review.
-
Mechanisms underlying anomalous diffusion in the plasma membrane.Curr Top Membr. 2015;75:167-207. doi: 10.1016/bs.ctm.2015.03.002. Epub 2015 Apr 15. Curr Top Membr. 2015. PMID: 26015283 Review.
Cited by
-
Active diffusion and advection in Drosophila oocytes result from the interplay of actin and microtubules.Nat Commun. 2017 Nov 15;8(1):1520. doi: 10.1038/s41467-017-01414-6. Nat Commun. 2017. PMID: 29142223 Free PMC article.
-
Endoplasmic reticulum network heterogeneity guides diffusive transport and kinetics.Biophys J. 2023 Aug 8;122(15):3191-3205. doi: 10.1016/j.bpj.2023.06.022. Epub 2023 Jul 3. Biophys J. 2023. PMID: 37401053 Free PMC article.
-
Intracellular transport dynamics revealed by single-particle tracking.Biophys Rep. 2021 Oct 31;7(5):413-427. doi: 10.52601/bpr.2021.210035. Biophys Rep. 2021. PMID: 37288105 Free PMC article.
-
Cytoplasmic Flow and Mixing Due to Deformation of Motile Cells.Biophys J. 2017 Nov 7;113(9):2077-2087. doi: 10.1016/j.bpj.2017.09.009. Biophys J. 2017. PMID: 29117530 Free PMC article.
-
Large-Scale, Wavelet-Based Analysis of Lysosomal Trajectories and Co-Movements of Lysosomes with Nanoparticle Cargos.Cells. 2022 Jan 13;11(2):270. doi: 10.3390/cells11020270. Cells. 2022. PMID: 35053385 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

