Microbial metabolomics in open microscale platforms
- PMID: 26842393
- PMCID: PMC4742997
- DOI: 10.1038/ncomms10610
Microbial metabolomics in open microscale platforms
Abstract
The microbial secondary metabolome encompasses great synthetic diversity, empowering microbes to tune their chemical responses to changing microenvironments. Traditional metabolomics methods are ill-equipped to probe a wide variety of environments or environmental dynamics. Here we introduce a class of microscale culture platforms to analyse chemical diversity of fungal and bacterial secondary metabolomes. By leveraging stable biphasic interfaces to integrate microculture with small molecule isolation via liquid-liquid extraction, we enable metabolomics-scale analysis using mass spectrometry. This platform facilitates exploration of culture microenvironments (including rare media typically inaccessible using established methods), unusual organic solvents for metabolite isolation and microbial mutants. Utilizing Aspergillus, a fungal genus known for its rich secondary metabolism, we characterize the effects of culture geometry and growth matrix on secondary metabolism, highlighting the potential use of microscale systems to unlock unknown or cryptic secondary metabolites for natural products discovery. Finally, we demonstrate the potential for this class of microfluidic systems to study interkingdom communication between fungi and bacteria.
Conflict of interest statement
D.J.B. has ownership in BellBrook Labs, LLC; Salus Discovery, LLC; Tasso, Inc.; and Stacks to the Future, LLC. E.B. has ownership in Salus Discovery, LLC; Tasso, Inc.; and Stacks to the Future, LLC. A.B.T. has ownership in Stacks to the Future, LLC. The remaining authors declare no competing financial interests.
Figures


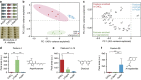


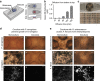
References
-
- Woloshuk C. P. & Shim W. B. Aflatoxins, fumonisins, and trichothecenes: a convergence of knowledge. FEMS Microbiol. Rev. 37, 94–109 (2012). - PubMed
-
- Friesen T. L., Faris J. D., Solomon P. S. & Oliver R. P. Host-specific toxins: effectors of necrotrophic pathogenicity. Cell Microbiol. 10, 1421–1428 (2008). - PubMed
-
- Brown S. H. et al. Oxygenase coordination is required for morphological transition and the host–fungus interaction of Aspergillus flavus. Mol. Plant Microbe Interact. 22, 882–894 (2009). - PubMed
-
- Wu F., Groopman J. D. & Pestka J. J. Public Health Impacts of Foodborne Mycotoxins. Annu. Rev. Food Sci. Technol. 5, 351–372 (2014). - PubMed
-
- Evidente A. et al. Fungal metabolites with anticancer activity. Nat. Prod. Rep. 31, 617 (2014). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

