Tankyrases Promote Homologous Recombination and Check Point Activation in Response to DSBs
- PMID: 26845027
- PMCID: PMC4741384
- DOI: 10.1371/journal.pgen.1005791
Tankyrases Promote Homologous Recombination and Check Point Activation in Response to DSBs
Abstract
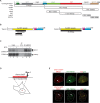
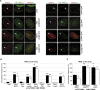
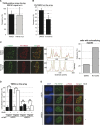
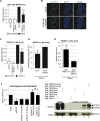
DNA lesions are sensed by a network of proteins that trigger the DNA damage response (DDR), a signaling cascade that acts to delay cell cycle progression and initiate DNA repair. The Mediator of DNA damage Checkpoint protein 1 (MDC1) is essential for spreading of the DDR signaling on chromatin surrounding Double Strand Breaks (DSBs) by acting as a scaffold for PI3K kinases and for ubiquitin ligases. MDC1 also plays a role both in Non-Homologous End Joining (NHEJ) and Homologous Recombination (HR) repair pathways. Here we identify two novel binding partners of MDC1, the poly (ADP-ribose) Polymerases (PARPs) TNKS1 and 2. We find that TNKSs are recruited to DNA lesions by MDC1 and regulate DNA end resection and BRCA1A complex stabilization at lesions leading to efficient DSB repair by HR and proper checkpoint activation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


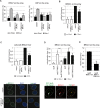
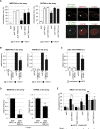
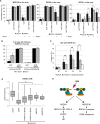
Similar articles
-
Ataxin-3 consolidates the MDC1-dependent DNA double-strand break response by counteracting the SUMO-targeted ubiquitin ligase RNF4.EMBO J. 2017 Apr 13;36(8):1066-1083. doi: 10.15252/embj.201695151. Epub 2017 Mar 8. EMBO J. 2017. PMID: 28275011 Free PMC article.
-
Remodeling and spacing factor 1 (RSF1) deposits centromere proteins at DNA double-strand breaks to promote non-homologous end-joining.Cell Cycle. 2013 Sep 15;12(18):3070-82. doi: 10.4161/cc.26033. Epub 2013 Aug 20. Cell Cycle. 2013. PMID: 23974106 Free PMC article.
-
REV7 counteracts DNA double-strand break resection and affects PARP inhibition.Nature. 2015 May 28;521(7553):541-544. doi: 10.1038/nature14328. Epub 2015 Mar 23. Nature. 2015. PMID: 25799992 Free PMC article.
-
The influence of heterochromatin on DNA double strand break repair: Getting the strong, silent type to relax.DNA Repair (Amst). 2010 Dec 10;9(12):1273-82. doi: 10.1016/j.dnarep.2010.09.013. Epub 2010 Oct 30. DNA Repair (Amst). 2010. PMID: 21036673 Review.
-
Regulation of repair pathway choice at two-ended DNA double-strand breaks.Mutat Res. 2017 Oct;803-805:51-55. doi: 10.1016/j.mrfmmm.2017.07.011. Epub 2017 Jul 29. Mutat Res. 2017. PMID: 28781144 Review.
Cited by
-
The PARP Enzyme Family and the Hallmarks of Cancer Part 1. Cell Intrinsic Hallmarks.Cancers (Basel). 2021 Apr 23;13(9):2042. doi: 10.3390/cancers13092042. Cancers (Basel). 2021. PMID: 33922595 Free PMC article. Review.
-
Fueling genome maintenance: On the versatile roles of NAD+ in preserving DNA integrity.J Biol Chem. 2022 Jun;298(6):102037. doi: 10.1016/j.jbc.2022.102037. Epub 2022 May 17. J Biol Chem. 2022. PMID: 35595095 Free PMC article. Review.
-
ADP-ribosylation signalling and human disease.Open Biol. 2019 Apr 26;9(4):190041. doi: 10.1098/rsob.190041. Open Biol. 2019. PMID: 30991935 Free PMC article. Review.
-
Functions of ADP-ribose transferases in the maintenance of telomere integrity.Cell Mol Life Sci. 2022 Mar 29;79(4):215. doi: 10.1007/s00018-022-04235-z. Cell Mol Life Sci. 2022. PMID: 35348914 Free PMC article. Review.
-
Identifying and Validating Tankyrase Binders and Substrates: A Candidate Approach.Methods Mol Biol. 2017;1608:445-473. doi: 10.1007/978-1-4939-6993-7_28. Methods Mol Biol. 2017. PMID: 28695526 Free PMC article.
References
-
- van Gent DC, Hoeijmakers JH, Kanaar R. Chromosomal stability and the DNA double-stranded break connection. Nat Rev Genet. 2001;2(3):196–206. Epub 2001/03/21. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

