Intra-Species Diversity and Panmictic Structure of Cryptosporidium parvum Populations in Cattle Farms in Northern Spain
- PMID: 26848837
- PMCID: PMC4746124
- DOI: 10.1371/journal.pone.0148811
Intra-Species Diversity and Panmictic Structure of Cryptosporidium parvum Populations in Cattle Farms in Northern Spain
Abstract
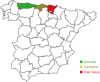
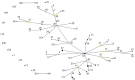
The intra-herd and intra-host genetic variability of 123 Cryptosporidium parvum isolates was investigated using a multilocus fragment typing approach with eleven variable-number tandem-repeat (VNTR) loci and the GP60 gene. Isolates were collected from intensively farmed diarrheic pre-weaned calves originating from 31 dairy farms in three adjoining regions in northern Spain (País Vasco, Cantabria and Asturias). The multilocus tool demonstrated an acceptable typeability, with 104/123 samples amplifying at all twelve loci. The ML2, TP14, GP60 and the previously un-described minisatellite at locus cgd2_3850 were the most discriminatory markers, while others may be dismissed as monomorphic (MSB) or less informative (CP47, ML1 and the novel minisatellites at loci Cgd1_3670 and Cgd6_3940). The 12-satellite typing tool provided a Hunter-Gaston index (HGDI) of 0.987 (95% CI, 0.982-0.992), and differentiated a total of 70 multilocus subtypes (MLTs). The inclusion of only the four most discriminatory markers dramatically reduced the number of MLTs (n: 44) but hardly reduced the HGDI value. A total of 54 MLTs were distinctive for individual farms, indicating that cryptosporidiosis is an endemic condition on most cattle farms. However, a high rate of mixed infections was detected, suggesting frequent meiotic recombination. Namely, multiple MLTs were seen in most farms where several specimens were analyzed (90.5%), with up to 9 MLTs being found on one farm, and individual specimens with mixed populations being reported on 11/29 farms. Bayesian Structure analysis showed that over 35% of isolates had mixed ancestry and analysis of evolutionary descent using the eBURST algorithm detected a high rate (21.4%) of MLTs appearing as singletons, indicating a high degree of genetic divergence. Linkage analysis found evidence of linkage equilibrium and an overall panmictic structure within the C. parvum population in this discrete geographical area.
Conflict of interest statement
Figures

Similar articles
-
Intra-Species Genetic Diversity and Clonal Structure of Cryptosporidium parvum in Sheep Farms in a Confined Geographical Area in Northeastern Spain.PLoS One. 2016 May 13;11(5):e0155336. doi: 10.1371/journal.pone.0155336. eCollection 2016. PLoS One. 2016. PMID: 27176718 Free PMC article.
-
Multilocus fragment analysis of Cryptosporidium parvum from pre-weaned calves in Colombia.Acta Trop. 2019 Apr;192:151-157. doi: 10.1016/j.actatropica.2019.02.005. Epub 2019 Feb 7. Acta Trop. 2019. PMID: 30738722
-
Multilocus fragment typing and genetic structure of Cryptosporidium parvum Isolates from diarrheic preweaned calves in Spain.Appl Environ Microbiol. 2011 Nov;77(21):7779-86. doi: 10.1128/AEM.00751-11. Epub 2011 Sep 9. Appl Environ Microbiol. 2011. PMID: 21908632 Free PMC article.
-
Host association of Cryptosporidium parvum populations infecting domestic ruminants in Spain.Appl Environ Microbiol. 2013 Sep;79(17):5363-71. doi: 10.1128/AEM.01168-13. Epub 2013 Jun 28. Appl Environ Microbiol. 2013. PMID: 23811515 Free PMC article.
-
Assessment of polymorphic genetic markers for multi-locus typing of Cryptosporidium parvum and Cryptosporidium hominis.Exp Parasitol. 2012 Oct;132(2):200-15. doi: 10.1016/j.exppara.2012.06.016. Epub 2012 Jul 7. Exp Parasitol. 2012. PMID: 22781277 Review.
Cited by
-
Direct Sequencing of Cryptosporidium in Stool Samples for Public Health.Front Public Health. 2019 Dec 11;7:360. doi: 10.3389/fpubh.2019.00360. eCollection 2019. Front Public Health. 2019. PMID: 31921734 Free PMC article. Review.
-
Molecular epidemiologic tools for waterborne pathogens Cryptosporidium spp. and Giardia duodenalis.Food Waterborne Parasitol. 2017 Sep 29;8-9:14-32. doi: 10.1016/j.fawpar.2017.09.002. eCollection 2017 Sep-Dec. Food Waterborne Parasitol. 2017. PMID: 32095639 Free PMC article. Review.
-
Intra-Species Genetic Diversity and Clonal Structure of Cryptosporidium parvum in Sheep Farms in a Confined Geographical Area in Northeastern Spain.PLoS One. 2016 May 13;11(5):e0155336. doi: 10.1371/journal.pone.0155336. eCollection 2016. PLoS One. 2016. PMID: 27176718 Free PMC article.
-
Occurrence and Molecular Characterization of Cryptosporidium spp. in Dairy Cattle and Dairy Buffalo in Yunnan Province, Southwest China.Animals (Basel). 2022 Apr 15;12(8):1031. doi: 10.3390/ani12081031. Animals (Basel). 2022. PMID: 35454277 Free PMC article.
-
Challenges for Cryptosporidium Population Studies.Genes (Basel). 2021 Jun 10;12(6):894. doi: 10.3390/genes12060894. Genes (Basel). 2021. PMID: 34200631 Free PMC article. Review.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

