Subcellular Fractionation Analysis of the Extraction of Ubiquitinated Polytopic Membrane Substrate during ER-Associated Degradation
- PMID: 26849222
- PMCID: PMC4743956
- DOI: 10.1371/journal.pone.0148327
Subcellular Fractionation Analysis of the Extraction of Ubiquitinated Polytopic Membrane Substrate during ER-Associated Degradation
Abstract
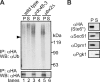
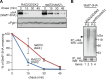
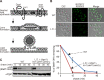
During ER-associated degradation (ERAD), misfolded polytopic membrane proteins are ubiquitinated and retrotranslocated to the cytosol for proteasomal degradation. However, our understanding as to how polytopic membrane proteins are extracted from the ER to the cytosol remains largely unclear. To better define the localization and physical properties of ubiquitinated polytopic membrane substrates in vivo, we performed subcellular fractionation analysis of Ste6*, a twelve transmembrane protein that is ubiquitinated primarily by Doa10 E3 ligase in yeast. Consistent with previous in vitro studies, ubiquitinated Ste6* was extracted from P20 (20,000 g pellet) fraction to S20 (20,000 g supernatant) fraction in a Cdc48/p97-dependent manner. Similarly, Ubx2p, which recruits Cdc48/p97 to the ER, facilitated the extraction of Ste6*. By contrast, lipid droplet formation, which was suggested to be dispensable for the degradation of Hrd1-substrates in yeast, was not required for the degradation of Ste6*. Intriguingly, we found that ubiquitinated Ste6* in the S20 fraction could be enriched by further centrifugation at 100,000 g. Although it is currently uncertain whether ubiquitinated Ste6* in P100 fraction is completely free from any lipids, membrane flotation analysis suggested the existence of two distinct populations of ubiquitinated Ste6* with different states of membrane association. Together, these results imply that ubiquitinated Ste6* may be sequestered into a putative quality control sub-structure by Cdc48/p97. Fractionation assays developed in the present study provide a means to further dissect the ill-defined post-ubiquitination step during ERAD of polytopic membrane substrates.
Conflict of interest statement
Figures


Similar articles
-
Endoplasmic Reticulum-associated Degradation of Pca1p, a Polytopic Protein, via Interaction with the Proteasome at the Membrane.J Biol Chem. 2016 Jul 15;291(29):15082-92. doi: 10.1074/jbc.M116.726265. Epub 2016 May 12. J Biol Chem. 2016. PMID: 27226596 Free PMC article.
-
A Cdc48 "Retrochaperone" Function Is Required for the Solubility of Retrotranslocated, Integral Membrane Endoplasmic Reticulum-associated Degradation (ERAD-M) Substrates.J Biol Chem. 2017 Feb 24;292(8):3112-3128. doi: 10.1074/jbc.M116.770610. Epub 2017 Jan 11. J Biol Chem. 2017. PMID: 28077573 Free PMC article.
-
Previously unknown role for the ubiquitin ligase Ubr1 in endoplasmic reticulum-associated protein degradation.Proc Natl Acad Sci U S A. 2013 Sep 17;110(38):15271-6. doi: 10.1073/pnas.1304928110. Epub 2013 Aug 29. Proc Natl Acad Sci U S A. 2013. PMID: 23988329 Free PMC article.
-
Mechanistic insights into ER-associated protein degradation.Curr Opin Cell Biol. 2018 Aug;53:22-28. doi: 10.1016/j.ceb.2018.04.004. Epub 2018 Apr 30. Curr Opin Cell Biol. 2018. PMID: 29719269 Free PMC article. Review.
-
Mitochondrial quality control by the ubiquitin-proteasome system.Biochem Soc Trans. 2011 Oct;39(5):1509-13. doi: 10.1042/BST0391509. Biochem Soc Trans. 2011. PMID: 21936843 Review.
Cited by
-
Potential Physiological Relevance of ERAD to the Biosynthesis of GPI-Anchored Proteins in Yeast.Int J Mol Sci. 2021 Jan 21;22(3):1061. doi: 10.3390/ijms22031061. Int J Mol Sci. 2021. PMID: 33494405 Free PMC article. Review.
-
The degradation-promoting roles of deubiquitinases Ubp6 and Ubp3 in cytosolic and ER protein quality control.PLoS One. 2020 May 13;15(5):e0232755. doi: 10.1371/journal.pone.0232755. eCollection 2020. PLoS One. 2020. PMID: 32401766 Free PMC article.
-
Hydroxyurea modulates thiol-disulfide homeostasis in the yeast endoplasmic reticulum.Life Sci Alliance. 2025 Jun 20;8(8):e202503225. doi: 10.26508/lsa.202503225. Print 2025 Aug. Life Sci Alliance. 2025. PMID: 40541417 Free PMC article.
-
The Dfm1 Derlin Is Required for ERAD Retrotranslocation of Integral Membrane Proteins.Mol Cell. 2018 Jan 18;69(2):306-320.e4. doi: 10.1016/j.molcel.2017.12.012. Mol Cell. 2018. PMID: 29351849 Free PMC article.
-
Protein Quality Control and Lipid Droplet Metabolism.Annu Rev Cell Dev Biol. 2020 Oct 6;36:115-139. doi: 10.1146/annurev-cellbio-031320-101827. Annu Rev Cell Dev Biol. 2020. PMID: 33021827 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

