The Type IV Pilus Assembly ATPase PilB of Myxococcus xanthus Interacts with the Inner Membrane Platform Protein PilC and the Nucleotide-binding Protein PilM
- PMID: 26851283
- PMCID: PMC4807279
- DOI: 10.1074/jbc.M115.701284
The Type IV Pilus Assembly ATPase PilB of Myxococcus xanthus Interacts with the Inner Membrane Platform Protein PilC and the Nucleotide-binding Protein PilM
Abstract
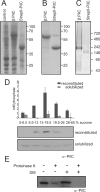
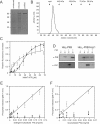
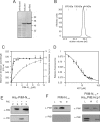
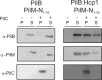
Type IV pili (T4P) are ubiquitous bacterial cell surface structures, involved in processes such as twitching motility, biofilm formation, bacteriophage infection, surface attachment, virulence, and natural transformation. T4P are assembled by machinery that can be divided into the outer membrane pore complex, the alignment complex that connects components in the inner and outer membrane, and the motor complex in the inner membrane and cytoplasm. Here, we characterize the inner membrane platform protein PilC, the cytosolic assembly ATPase PilB of the motor complex, and the cytosolic nucleotide-binding protein PilM of the alignment complex of the T4P machinery ofMyxococcus xanthus PilC was purified as a dimer and reconstituted into liposomes. PilB was isolated as a monomer and bound ATP in a non-cooperative manner, but PilB fused to Hcp1 ofPseudomonas aeruginosaformed a hexamer and bound ATP in a cooperative manner. Hexameric but not monomeric PilB bound to PilC reconstituted in liposomes, and this binding stimulated PilB ATPase activity. PilM could only be purified when it was stabilized by a fusion with a peptide corresponding to the first 16 amino acids of PilN, supporting an interaction between PilM and PilN(1-16). PilM-N(1-16) was isolated as a monomer that bound but did not hydrolyze ATP. PilM interacted directly with PilB, but only with PilC in the presence of PilB, suggesting an indirect interaction. We propose that PilB interacts with PilC and with PilM, thus establishing the connection between the alignment and the motor complex.
Keywords: ATPase; Myxococcus xanthus; cell motility; membrane reconstitution; membrane transport; secretion; type IV pili.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures
Similar articles
-
PilN Binding Modulates the Structure and Binding Partners of the Pseudomonas aeruginosa Type IVa Pilus Protein PilM.J Biol Chem. 2016 May 20;291(21):11003-15. doi: 10.1074/jbc.M116.718353. Epub 2016 Mar 28. J Biol Chem. 2016. PMID: 27022027 Free PMC article.
-
Outside-in assembly pathway of the type IV pilus system in Myxococcus xanthus.J Bacteriol. 2014 Jan;196(2):378-90. doi: 10.1128/JB.01094-13. Epub 2013 Nov 1. J Bacteriol. 2014. PMID: 24187092 Free PMC article.
-
The platform protein is essential for type IV pilus biogenesis.J Biol Chem. 2013 Apr 5;288(14):9721-9728. doi: 10.1074/jbc.M113.453506. Epub 2013 Feb 14. J Biol Chem. 2013. PMID: 23413032 Free PMC article.
-
Biogenesis of Pseudomonas aeruginosa type IV pili and regulation of their function.Environ Microbiol. 2015 Nov;17(11):4148-63. doi: 10.1111/1462-2920.12849. Epub 2015 Jun 25. Environ Microbiol. 2015. PMID: 25808785 Review.
-
Pulling together with type IV pili.J Mol Microbiol Biotechnol. 2004;7(1-2):52-62. doi: 10.1159/000077869. J Mol Microbiol Biotechnol. 2004. PMID: 15170403 Review.
Cited by
-
Data-Driven Models Reveal Mutant Cell Behaviors Important for Myxobacterial Aggregation.mSystems. 2020 Jul 14;5(4):e00518-20. doi: 10.1128/mSystems.00518-20. mSystems. 2020. PMID: 32665330 Free PMC article.
-
The type IV pilus protein PilU functions as a PilT-dependent retraction ATPase.PLoS Genet. 2019 Sep 16;15(9):e1008393. doi: 10.1371/journal.pgen.1008393. eCollection 2019 Sep. PLoS Genet. 2019. PMID: 31525185 Free PMC article.
-
Tight-packing of large pilin subunits provides distinct structural and mechanical properties for the Myxococcus xanthus type IVa pilus.Proc Natl Acad Sci U S A. 2024 Apr 23;121(17):e2321989121. doi: 10.1073/pnas.2321989121. Epub 2024 Apr 16. Proc Natl Acad Sci U S A. 2024. PMID: 38625941 Free PMC article.
-
Zinc coordination is essential for the function and activity of the type II secretion ATPase EpsE.Microbiologyopen. 2016 Oct;5(5):870-882. doi: 10.1002/mbo3.376. Epub 2016 May 10. Microbiologyopen. 2016. PMID: 27168165 Free PMC article.
-
The molecular mechanism of the type IVa pilus motors.Nat Commun. 2017 May 5;8:15091. doi: 10.1038/ncomms15091. Nat Commun. 2017. PMID: 28474682 Free PMC article.
References
-
- Pohlschroder M., Ghosh A., Tripepi M., and Albers S.-V. (2011) Archaeal type IV pilus-like structures—evolutionarily conserved prokaryotic surface organelles. Curr. Opin. Microbiol. 14, 357–363 - PubMed
-
- Ayers M., Howell P. L., and Burrows L. L. (2010) Architecture of the type II secretion and type IV pilus machineries. Future Microbiol. 5, 1203–1218 - PubMed
-
- Hobbs M., and Mattick J. S. (1993) Common components in the assembly of type 4 fimbriae, DNA transfer systems, filamentous phage and protein-secretion apparatus: a general system for the formation of surface-associated protein complexes. Mol. Microbiol. 10, 233–243 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

