Truncating Erythropoietin Receptor Rearrangements in Acute Lymphoblastic Leukemia
- PMID: 26859458
- PMCID: PMC4750652
- DOI: 10.1016/j.ccell.2015.12.013
Truncating Erythropoietin Receptor Rearrangements in Acute Lymphoblastic Leukemia
Abstract
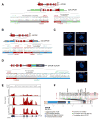
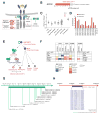
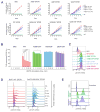
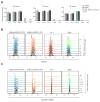
Chromosomal rearrangements are a hallmark of acute lymphoblastic leukemia (ALL) and are important ALL initiating events. We describe four different rearrangements of the erythropoietin receptor gene EPOR in Philadelphia chromosome-like (Ph-like) ALL. All of these rearrangements result in truncation of the cytoplasmic tail of EPOR at residues similar to those mutated in primary familial congenital polycythemia, with preservation of the proximal tyrosine essential for receptor activation and loss of distal regulatory residues. This resulted in deregulated EPOR expression, hypersensitivity to erythropoietin stimulation, and heightened JAK-STAT activation. Expression of truncated EPOR in mouse B cell progenitors induced ALL in vivo. Human leukemic cells with EPOR rearrangements were sensitive to JAK-STAT inhibition, suggesting a therapeutic option in high-risk ALL.
Copyright © 2016 Elsevier Inc. All rights reserved.
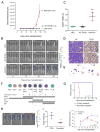
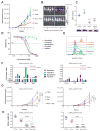
Figures

Similar articles
-
Erythropoietin hypersensitivity in primary familial and congenital polycythemia: role of tyrosines Y285 and Y344 in erythropoietin receptor cytoplasmic domain.Biochim Biophys Acta. 2005 Apr 15;1740(1):17-28. doi: 10.1016/j.bbadis.2005.03.003. Epub 2005 Apr 1. Biochim Biophys Acta. 2005. PMID: 15878737
-
Development of acute lymphoblastic leukemia with IgH-EPOR in a patient with secondary erythrocytosis.Int J Hematol. 2016 Dec;104(6):741-743. doi: 10.1007/s12185-016-2083-2. Epub 2016 Aug 20. Int J Hematol. 2016. PMID: 27544511
-
Primary familial polycythemia: a frameshift mutation in the erythropoietin receptor gene and increased sensitivity of erythroid progenitors to erythropoietin.Blood. 1995 Jul 1;86(1):15-22. Blood. 1995. PMID: 7795221
-
The biology of Philadelphia chromosome-like ALL.Best Pract Res Clin Haematol. 2017 Sep;30(3):212-221. doi: 10.1016/j.beha.2017.07.003. Epub 2017 Jul 6. Best Pract Res Clin Haematol. 2017. PMID: 29050694 Review.
-
"Benign erythrocytosis" and other familial and congenital polycythemias.Eur J Haematol. 1996 Oct;57(4):263-8. Eur J Haematol. 1996. PMID: 8982288 Review.
Cited by
-
Recent advances and novel treatment paradigms in acute lymphocytic leukemia.Ther Adv Hematol. 2016 Oct;7(5):252-269. doi: 10.1177/2040620716652289. Epub 2016 Jun 29. Ther Adv Hematol. 2016. PMID: 27695616 Free PMC article. Review.
-
Treatment of Young Adults with Acute Lymphoblastic Leukemia.Curr Hematol Malig Rep. 2017 Jun;12(3):187-196. doi: 10.1007/s11899-017-0377-y. Curr Hematol Malig Rep. 2017. PMID: 28353017 Review.
-
Philadelphia chromosome-like acute lymphoblastic leukemia.Blood. 2017 Nov 9;130(19):2064-2072. doi: 10.1182/blood-2017-06-743252. Epub 2017 Oct 2. Blood. 2017. PMID: 28972016 Free PMC article. Review.
-
Ph-like acute lymphoblastic leukemia.Hematology Am Soc Hematol Educ Program. 2016 Dec 2;2016(1):561-566. doi: 10.1182/asheducation-2016.1.561. Hematology Am Soc Hematol Educ Program. 2016. PMID: 27913529 Free PMC article. Review.
-
B-ALL Complexity: Is Targeted Therapy Still A Valuable Approach for Pediatric Patients?Cancers (Basel). 2020 Nov 24;12(12):3498. doi: 10.3390/cancers12123498. Cancers (Basel). 2020. PMID: 33255367 Free PMC article. Review.
References
-
- Alpar D, Wren D, Ermini L, Mansur MB, van Delft FW, Bateman CM, Titley I, Kearney L, Szczepanski T, Gonzalez D, et al. Clonal origins of ETV6-RUNX1(+) acute lymphoblastic leukemia: studies in monozygotic twins. Leukemia. 2015;29:839–846. - PubMed
-
- Bento C, Percy MJ, Gardie B, Maia TM, van Wijk R, Perrotta S, Della Ragione F, Almeida H, Rossi C, Girodon F, et al. Genetic basis of congenital erythrocytosis: mutation update and online databases. Hum Mutat. 2014;35:15–26. - PubMed
-
- Den Boer ML, van Slegtenhorst M, De Menezes RX, Cheok MH, Buijs-Gladdines JG, Peters ST, Van Zutven LJ, Beverloo HB, Van der Spek PJ, Escherich G, et al. A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol. 2009;10:125–134. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U24 CA196173/CA/NCI NIH HHS/United States
- U10 CA98413/CA/NCI NIH HHS/United States
- U10 CA021115/CA/NCI NIH HHS/United States
- CA21765/CA/NCI NIH HHS/United States
- U10 CA101140/CA/NCI NIH HHS/United States
- P30 CA021765/CA/NCI NIH HHS/United States
- CA180827/CA/NCI NIH HHS/United States
- U01 CA157937/CA/NCI NIH HHS/United States
- U10 CA098543/CA/NCI NIH HHS/United States
- U10 CA180794/CA/NCI NIH HHS/United States
- U24 CA114737/CA/NCI NIH HHS/United States
- U10 CA98543/CA/NCI NIH HHS/United States
- U24 CA196172/CA/NCI NIH HHS/United States
- CA145707/CA/NCI NIH HHS/United States
- U10 CA098413/CA/NCI NIH HHS/United States
- U24 CA114766/CA/NCI NIH HHS/United States
- P30 CA008748/CA/NCI NIH HHS/United States
- U10 CA180791/CA/NCI NIH HHS/United States
- RC1 CA145707/CA/NCI NIH HHS/United States
- U10 CA180820/CA/NCI NIH HHS/United States
- U10 CA180827/CA/NCI NIH HHS/United States
- P50 GM115279/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

