Can MLVA Differentiate among Endemic-Like MRSA Isolates with Identical Spa-Type in a Low-Prevalence Region?
- PMID: 26859765
- PMCID: PMC4747572
- DOI: 10.1371/journal.pone.0148772
Can MLVA Differentiate among Endemic-Like MRSA Isolates with Identical Spa-Type in a Low-Prevalence Region?
Abstract
The prevalence of methicillin-resistant Staphylococcus aureus (MRSA) in Norway is low, but an endemic-like MRSA clone with Staphylococcal protein A (spa)-type t304 has been established especially in nursing homes in the Oslo region causing several large outbreaks. The challenge was that spa-typing and the gold standard Pulsed-Field Gel Electrophoresis (PFGE) were inadequate in discriminating isolates in outbreak investigations. Additional higher resolution genotyping methods were needed. The aims of this study were a) to evaluate whether Multiple-Locus Variable number of tandem repeat Analysis (MLVA) could differentiate within the PFGE clusters between epidemiologically related and unrelated endemic-like ST8-MRSA-IV-t304-PVL-neg (MRSA-t304) isolates and b) investigate the evolution of the endemic-like MRSA-t304 clone over a 15-year time period. All MRSA-t304 isolates detected in the region from 1998 through April 2013 were included. In total, 194 of 197 isolates were available for PFGE and MLVA analyses. PFGE results on isolates from 1998-2010 have been published previously. Two PFGE clusters subdivided into eight MLVA types were detected. One major outbreak clone (PFGE cluster C2/ MLVA type MT5045) appeared from 2004 to 2011 causing long-lasting and large outbreaks in seven nursing homes and one hospital. Five new MLVA types (N = 9 isolates) differing in only one VNTR compared to the outbreak clone C2/MT5045 were detected, but only one (C2/MT5044) was seen after 2011. We suggest that MLVA can replace PFGE analysis, but MLVA may not be the optimal method in this setting as it did not discriminate between all epidemiologically unrelated isolates. The results may indicate that all eight outbreaks in different locations within the PFGE C2 cluster may be branches of one large regional outbreak. The major outbreak strain C2/MT5045 may now, however, be under control, extinguished or has moved geographically.
Conflict of interest statement
Figures
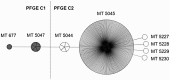
References
-
- National guidelines for the prevention of spread of methicillin-resistant Staphylococcus aureus (MRSA) in health institutions. National Insitute of Public Health and the Norwegian Directorate of Health. 2009.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

