Current Experimental Methods for Characterizing Protein-Protein Interactions
- PMID: 26864455
- PMCID: PMC7162211
- DOI: 10.1002/cmdc.201500495
Current Experimental Methods for Characterizing Protein-Protein Interactions
Abstract
Protein molecules often interact with other partner protein molecules in order to execute their vital functions in living organisms. Characterization of protein-protein interactions thus plays a central role in understanding the molecular mechanism of relevant protein molecules, elucidating the cellular processes and pathways relevant to health or disease for drug discovery, and charting large-scale interaction networks in systems biology research. A whole spectrum of methods, based on biophysical, biochemical, or genetic principles, have been developed to detect the time, space, and functional relevance of protein-protein interactions at various degrees of affinity and specificity. This article presents an overview of these experimental methods, outlining the principles, strengths and limitations, and recent developments of each type of method.
Keywords: biochemical methods; biophysical methods; genetic methods; protein-protein interactions.
© 2016 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures


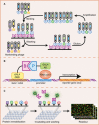
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

