Hepatitis C virus NS3-4A inhibits the peroxisomal MAVS-dependent antiviral signalling response
- PMID: 26865163
- PMCID: PMC5125814
- DOI: 10.1111/jcmm.12801
Hepatitis C virus NS3-4A inhibits the peroxisomal MAVS-dependent antiviral signalling response
Abstract
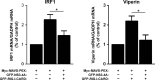
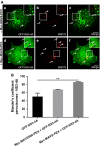
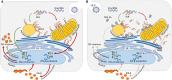
Hepatitis C virus (HCV) is the cause of one of the most prevalent viral infections worldwide. Upon infection, the HCV genome activates the RIG-I-MAVS signalling pathway leading to the production of direct antiviral effectors which prevent important steps in viral propagation. MAVS localizes at peroxisomes and mitochondria and coordinate the activation of an effective antiviral response: peroxisomal MAVS is responsible for a rapid but short-termed antiviral response, while the mitochondrial MAVS is associated with the activation of a stable response with delayed kinetics. The HCV NS3-4A protease was shown to specifically cleave the mitochondrial MAVS, inhibiting the downstream response. In this study, we have analysed whether HCV NS3-4A is also able to cleave the peroxisomal MAVS and whether this would have any effect on the cellular antiviral response. We show that NS3-4A is indeed able to specifically cleave this protein and release it into the cytosol, a mechanism that seems to occur at a similar kinetic rate as the cleavage of the mitochondrial MAVS. Under these conditions, RIG-I-like receptor (RLR) signalling from peroxisomes is blocked and antiviral gene expression is inhibited. Our results also show that NS3-4A is able to localize at peroxisomes in the absence of MAVS. However, mutation studies have shown that this localization pattern is preferred in the presence of a fully cleavable MAVS. These findings present evidence of a viral evasion strategy that disrupts RLR signalling on peroxisomes and provide an excellent example of how a single viral evasion strategy can block innate immune signalling from different organelles.
Keywords: Hepatitis C virus; MAVS; NS3-4A; peroxisomes.
© 2016 The Authors. Journal of Cellular and Molecular Medicine published by John Wiley & Sons Ltd and Foundation for Cellular and Molecular Medicine.
Figures


Similar articles
-
Recruitment of an interferon molecular signaling complex to the mitochondrial membrane: disruption by hepatitis C virus NS3-4A protease.Biochem Pharmacol. 2006 Nov 30;72(11):1477-84. doi: 10.1016/j.bcp.2006.06.030. Epub 2006 Jul 31. Biochem Pharmacol. 2006. PMID: 16876765
-
Activation of Type I and III Interferon Response by Mitochondrial and Peroxisomal MAVS and Inhibition by Hepatitis C Virus.PLoS Pathog. 2015 Nov 20;11(11):e1005264. doi: 10.1371/journal.ppat.1005264. eCollection 2015 Nov. PLoS Pathog. 2015. PMID: 26588843 Free PMC article.
-
Hepacivirus NS3/4A Proteases Interfere with MAVS Signaling in both Their Cognate Animal Hosts and Humans: Implications for Zoonotic Transmission.J Virol. 2016 Nov 14;90(23):10670-10681. doi: 10.1128/JVI.01634-16. Print 2016 Dec 1. J Virol. 2016. PMID: 27654291 Free PMC article.
-
[Chronic hepatitis C virus infection attenuates host antiviral innate immune response].Nihon Rinsho. 2015 Feb;73(2):234-8. Nihon Rinsho. 2015. PMID: 25764676 Review. Japanese.
-
Orchestrating the interferon antiviral response through the mitochondrial antiviral signaling (MAVS) adapter.Curr Opin Immunol. 2011 Oct;23(5):564-72. doi: 10.1016/j.coi.2011.08.001. Epub 2011 Aug 22. Curr Opin Immunol. 2011. PMID: 21865020 Review.
Cited by
-
Peroxisomes are platforms for cytomegalovirus' evasion from the cellular immune response.Sci Rep. 2016 May 16;6:26028. doi: 10.1038/srep26028. Sci Rep. 2016. PMID: 27181750 Free PMC article.
-
Infection of Glia by Human Pegivirus Suppresses Peroxisomal and Antiviral Signaling Pathways.J Virol. 2021 Nov 9;95(23):e0107421. doi: 10.1128/JVI.01074-21. Epub 2021 Sep 15. J Virol. 2021. PMID: 34524914 Free PMC article.
-
Peroxisomes in Immune Response and Inflammation.Int J Mol Sci. 2019 Aug 8;20(16):3877. doi: 10.3390/ijms20163877. Int J Mol Sci. 2019. PMID: 31398943 Free PMC article. Review.
-
Host mitochondria: more than an organelle in SARS-CoV-2 infection.Front Cell Infect Microbiol. 2023 Aug 25;13:1228275. doi: 10.3389/fcimb.2023.1228275. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37692170 Free PMC article. Review.
-
Mitochondria and Mitochondrial DNA: Key Elements in the Pathogenesis and Exacerbation of the Inflammatory State Caused by COVID-19.Medicina (Kaunas). 2021 Sep 3;57(9):928. doi: 10.3390/medicina57090928. Medicina (Kaunas). 2021. PMID: 34577851 Free PMC article. Review.
References
-
- Chevaliez S, Pawlotsky J‐M. HCV Genome and Life Cycle In: Tan S, editor. Hepatitis C Viruses Genomes and Molecular Biology. Norfolk (UK): Horizon Bioscience; 2006. pp. 5–47. - PubMed
-
- Sarrazin C, Hézode C, Zeuzem S, et al Antiviral strategies in hepatitis C virus infection. J Hepatol. 2012; 56: S88–100. - PubMed
-
- Yoneyama M, Kikuchi M, Natsukawa T, et al The RNA helicase RIG‐I has an essential function in double‐stranded RNA‐induced innate antiviral responses. Nat Immunol. 2004; 5: 730–7. - PubMed
-
- Cao X, Ding Q, Lu J, et al MDA5 plays a critical role in interferon response during hepatitis C virus infection. J Hepatol. 2014; 62: 771–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

