Human Neuropeptide S Receptor Is Activated via a Gαq Protein-biased Signaling Cascade by a Human Neuropeptide S Analog Lacking the C-terminal 10 Residues
- PMID: 26865629
- PMCID: PMC4817180
- DOI: 10.1074/jbc.M115.704122
Human Neuropeptide S Receptor Is Activated via a Gαq Protein-biased Signaling Cascade by a Human Neuropeptide S Analog Lacking the C-terminal 10 Residues
Abstract
Human neuropeptide S (NPS) and its cognate receptor regulate important biological functions in the brain and have emerged as a future therapeutic target for treatment of a variety of neurological and psychiatric diseases. The human NPS (hNPS) receptor has been shown to dually couple to Gαs- and Gαq-dependent signaling pathways. The human NPS analog hNPS-(1-10), lacking 10 residues from the C terminus, has been shown to stimulate Ca(2+)mobilization in a manner comparable with full-length hNPSin vitrobut seems to fail to induce biological activityin vivo Here, results derived from a number of cell-based functional assays, including intracellular cAMP-response element (CRE)-driven luciferase activity, Ca(2+)mobilization, and ERK1/2 phosphorylation, show that hNPS-(1-10) preferentially activates Gαq-dependent Ca(2+)mobilization while exhibiting less activity in triggering Gαs-dependent CRE-driven luciferase activity. We further demonstrate that both Gαq- and Gαs-coupled signaling pathways contribute to full-length hNPS-mediated activation of ERK1/2, whereas hNPS-(1-10)-promoted ERK1/2 activation is completely inhibited by the Gαqinhibitor UBO-QIC but not by the PKA inhibitor H89. Moreover, the results of Ala-scanning mutagenesis of hNPS-(1-13) indicated that residues Lys(11)and Lys(12)are structurally crucial for the hNPS receptor to couple to Gαs-dependent signaling. In conclusion, our findings demonstrate that hNPS-(1-10) is a biased agonist favoring Gαq-dependent signaling. It may represent a valuable chemical probe for further investigation of the therapeutic potential of human NPS receptor-directed signalingin vivo.
Keywords: G protein; G protein-coupled receptor (GPCR); extracellular-signal-regulated kinase (ERK); neuropeptide; signal transduction.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures
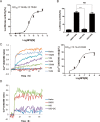


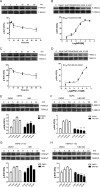
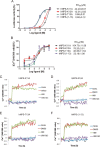
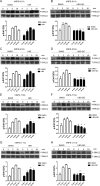
Similar articles
-
Specific activation of the G protein-coupled receptor BNGR-A21 by the neuropeptide corazonin from the silkworm, Bombyx mori, dually couples to the G(q) and G(s) signaling cascades.J Biol Chem. 2013 Apr 26;288(17):11662-75. doi: 10.1074/jbc.M112.441675. Epub 2013 Mar 1. J Biol Chem. 2013. PMID: 23457297 Free PMC article.
-
CAPA periviscerokinin-mediated activation of MAPK/ERK signaling through Gq-PLC-PKC-dependent cascade and reciprocal ERK activation-dependent internalized kinetics of Bom-CAPA-PVK receptor 2.Insect Biochem Mol Biol. 2018 Jul;98:1-15. doi: 10.1016/j.ibmb.2018.04.007. Epub 2018 May 4. Insect Biochem Mol Biol. 2018. PMID: 29730398
-
Targeted Elimination of G Proteins and Arrestins Defines Their Specific Contributions to Both Intensity and Duration of G Protein-coupled Receptor Signaling.J Biol Chem. 2016 Dec 30;291(53):27147-27159. doi: 10.1074/jbc.M116.754887. Epub 2016 Nov 16. J Biol Chem. 2016. PMID: 27852822 Free PMC article.
-
Gaq proteins: molecular pharmacology and therapeutic potential.Cell Mol Life Sci. 2017 Apr;74(8):1379-1390. doi: 10.1007/s00018-016-2405-9. Epub 2016 Nov 4. Cell Mol Life Sci. 2017. PMID: 27815595 Free PMC article. Review.
-
Gαq signalling from endosomes: A new conundrum.Br J Pharmacol. 2025 Jul;182(14):3068-3089. doi: 10.1111/bph.16248. Epub 2023 Oct 16. Br J Pharmacol. 2025. PMID: 37740273 Review.
Cited by
-
Macrocyclic Gq Protein Inhibitors FR900359 and/or YM-254890-Fit for Translation?ACS Pharmacol Transl Sci. 2021 Feb 19;4(2):888-897. doi: 10.1021/acsptsci.1c00021. eCollection 2021 Apr 9. ACS Pharmacol Transl Sci. 2021. PMID: 33860209 Free PMC article.
-
Selective G protein signaling driven by substance P-neurokinin receptor dynamics.Nat Chem Biol. 2022 Jan;18(1):109-115. doi: 10.1038/s41589-021-00890-8. Epub 2021 Oct 28. Nat Chem Biol. 2022. PMID: 34711980 Free PMC article.
-
Identification of the first biased NPS receptor agonist that retains anxiolytic and memory promoting effects with reduced levels of locomotor stimulation.Neuropharmacology. 2017 May 15;118:69-78. doi: 10.1016/j.neuropharm.2017.03.001. Epub 2017 Mar 3. Neuropharmacology. 2017. PMID: 28267583 Free PMC article.
-
An experimental strategy to probe Gq contribution to signal transduction in living cells.J Biol Chem. 2021 Jan-Jun;296:100472. doi: 10.1016/j.jbc.2021.100472. Epub 2021 Feb 25. J Biol Chem. 2021. PMID: 33639168 Free PMC article.
-
Heterotrimeric Gq proteins as therapeutic targets?J Biol Chem. 2020 Apr 17;295(16):5206-5215. doi: 10.1074/jbc.REV119.007061. Epub 2020 Mar 2. J Biol Chem. 2020. PMID: 32122969 Free PMC article. Review.
References
-
- Xu Y. L., Reinscheid R. K., Huitron-Resendiz S., Clark S. D., Wang Z., Lin S. H., Brucher F. A., Zeng J., Ly N. K., Henriksen S. J., de Lecea L., and Civelli O. (2004) Neuropeptide S: a neuropeptide promoting arousal and anxiolytic-like effects. Neuron 43, 487–497 - PubMed
-
- Sato S S. Y., Miyajima N, Yoshimura K. (September 19, 2002) Novel G protein-coupled receptor protein and DNA thereof. International Patent Application WO2002072829 A1
-
- Xu Y. L., Gall C. M., Jackson V. R., Civelli O., and Reinscheid R. K. (2007) Distribution of neuropeptide S receptor mRNA and neurochemical characteristics of neuropeptide S-expressing neurons in the rat brain. J. Comp. Neurol. 500, 84–102 - PubMed
-
- Leonard S. K., and Ring R. H. (2011) Immunohistochemical localization of the neuropeptide S receptor in the rat central nervous system. Neuroscience 172, 153–163 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

